
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 21,833,627 – 21,833,728 |
| Length | 101 |
| Max. P | 0.826512 |

| Location | 21,833,627 – 21,833,728 |
|---|---|
| Length | 101 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 57.92 |
| Shannon entropy | 0.87901 |
| G+C content | 0.47025 |
| Mean single sequence MFE | -20.32 |
| Consensus MFE | -5.69 |
| Energy contribution | -5.70 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.28 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.826512 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 21833627 101 - 24543557 AAGAAC----UACAUACGUAUGUACUUCUGCC-CACAUCCCCUUCAGCCACCUUCUCUUCAUCUUU-----CUGGUUUCGGCAAUGGGAAAUUUCCGUAGCCACCGCACAG--- .((((.----(((((....))))).))))((.-............(((((................-----.)))))..(((.((((((...)))))).)))...))....--- ( -19.23, z-score = -1.44, R) >droVir3.scaffold_13049 12866920 90 + 25233164 --------------AAGAACUCUCAGCGCACCACACUGUUCUCCUACCCACC-CCUUCUAGUGGAC----AAG--UUUCGGCAAUUGGAAAUUUCCACAGCCGCAACACUG--- --------------.(((((.................))))).....((((.-.......))))..----..(--((.((((...((((....))))..)))).)))....--- ( -19.33, z-score = -1.67, R) >droWil1.scaffold_180949 4325820 106 + 6375548 ----AAGAACUACCUUUCUCUAUCCCUGUCCUAACCCCUCCCCCCACCAACCGUCCUUUAAUGUAU----AAAGUUUUCGGCAAUAGGAAAUUUCCGUUGACGUUGGCAUACAG ----.((((......))))......((((.(((((...............(((..(((((.....)----))))....)))((((.(((....)))))))..)))))...)))) ( -14.00, z-score = -0.03, R) >droPer1.super_9 50950 105 + 3637205 AAGAAC----UAUAUG--UACAUACCUUUGGUAUAAGCUCCACAUAUGCACAUACUACACACGUAUAUCGGUUUGUUUCGGCAUUAGGAAAUUUCCGUAGCCAGCGCACAG--- ..((((----(..(((--((((((((...)))))..((.........)).............)))))).))))).....(((....(((....)))...))).........--- ( -18.80, z-score = 0.09, R) >dp4.chrXR_group6 8038600 105 + 13314419 AAGAAC----UAUAUG--UACAUACCUUUAGUAUAAGCUCCACAUAGGCACAUACUACACACGUAUAUCGGUUUGUUUCGGCAUUCGGAAAUUUCCGUAGCCAGCGCACAG--- ..((((----(..(((--(((.......((((((..(((.......)))..)))))).....)))))).))))).....(((...((((....))))..))).........--- ( -21.60, z-score = -1.37, R) >droAna3.scaffold_13337 20936696 83 - 23293914 -------------------AAGAACUACCUUUUCCGAUCCCCUACGA-CGCUUCCCG---AUGCUU-----CUCGUUUCGGCAAUAGGAAAUUUCCGUAGCUACUGCACAG--- -------------------........(((.((((((......((((-.((......---..))..-----.)))).)))).)).)))........((.((....))))..--- ( -14.10, z-score = -0.37, R) >droEre2.scaffold_4784 21506273 103 - 25762168 AAGAACUACGUACAUACGUAUGUACUUCUGCU-GAGACCCCCUUCGGCCCCCUUCUCA--GUCUUU-----CUGGUUUCGGCAAUGGGAAAUUUCCGUAGCCACCGCACAG--- .((((.(((((((....))))))).))))(((-((((....(....)......)))))--))....-----(((((...(((.((((((...)))))).)))...)).)))--- ( -30.20, z-score = -2.68, R) >droYak2.chr3L 22099343 103 - 24197627 AAGAAC----UAUAUA--CAUACGUAUGUACU-UCUGCUCAGGUCCACUUCAGUCCCCUUCUCUGUGUU-UCUGGUUUCGGCAAUGGGAAAUUUCCGUAGCCCCCGCACAG--- .((((.----((((((--(....))))))).)-)))(((((((..(((...((.......))..)))..-)))))....(((.((((((...)))))).)))...))....--- ( -27.10, z-score = -2.43, R) >droSec1.super_11 1754740 101 - 2888827 AAGAACUACAUACGUA--UAUGUACUUCUGCU-CACAUCCCCUUCAACCCCCUUCUCC--AUCUUU-----CUGAUUUCGGCAAUGGGAAAUUUCCGUAGCCACCGCACAG--- .((((.((((((....--)))))).))))((.-.........................--(((...-----..)))...(((.((((((...)))))).)))...))....--- ( -18.80, z-score = -2.46, R) >droSim1.chr3L_random 925229 101 - 1049610 AAGAACUACAUACGUA--UAUGUACUUCUGCU-CACAUCCCCUUCAACCCCCUUCUCC--AUCUUU-----CUGGUUUCGGCAAUGGGAAAUUUCCGUAGCCACCGCACAG--- .((((.((((((....--)))))).))))((.-.......................((--(.....-----.)))....(((.((((((...)))))).)))...))....--- ( -20.00, z-score = -2.22, R) >consensus AAGAAC____UACAUA__UAUGUACUUCUGCU_CACACCCCCUUCAACCACCUUCUCC__AUCUUU_____CUGGUUUCGGCAAUGGGAAAUUUCCGUAGCCACCGCACAG___ ...............................................................................(((....(((....)))...)))............ ( -5.69 = -5.70 + 0.01)

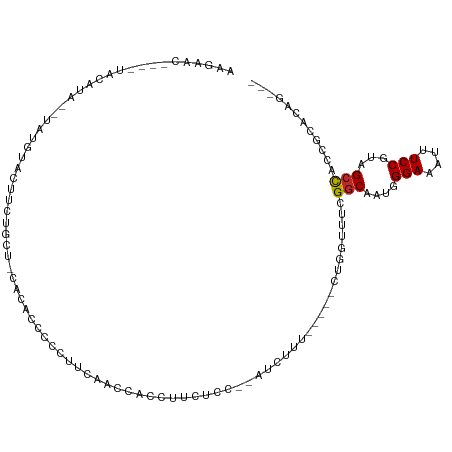
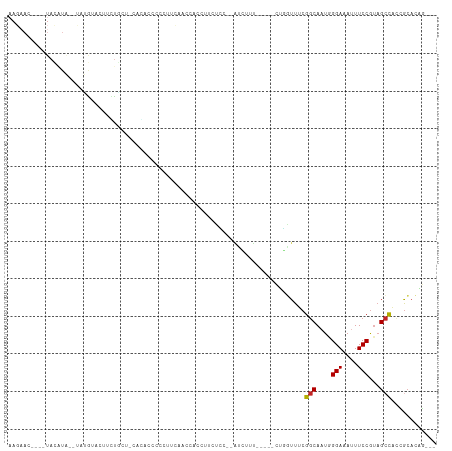
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:43:04 2011