
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 21,786,366 – 21,786,523 |
| Length | 157 |
| Max. P | 0.792930 |

| Location | 21,786,366 – 21,786,523 |
|---|---|
| Length | 157 |
| Sequences | 9 |
| Columns | 170 |
| Reading direction | reverse |
| Mean pairwise identity | 83.80 |
| Shannon entropy | 0.32820 |
| G+C content | 0.43778 |
| Mean single sequence MFE | -47.40 |
| Consensus MFE | -32.84 |
| Energy contribution | -32.42 |
| Covariance contribution | -0.41 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.792930 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 21786366 157 - 24543557 GCUAUGAAAUUAUGGCACCGUGCCGCCAUAUUCGGUUGUCAGGGUUUCGUUAUUGAGU--GGCGACAUAAAUUAAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGGACCAUAAACUC-CUUGACAAGUC-AACAGUA---GCCACUCCAGCGAGGAGUAGGAAA------ (((((....((((((..((((.(((((.(((((.(((((((....((((....)))))--)))))).........((((........)))))))))))))).)))).))))))....-.((((....))-))..)))---)).(((((.....)))))......------ ( -51.00, z-score = -2.37, R) >droSim1.chr3L 21129435 157 - 22553184 GCUAUGAAAUUAUGGCACCGUGCCGCCAUAUUCGGUUGUCAGGGUUUCGUUAUUGAGU--GGCGACAUAAAUUAAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGGACCAUAAACUC-CUUGACAAGUC-AACAGUA---GCCACUCCAGUGAGAAGUUGGAAA------ ((((((....))))))(((.(((.(((......))).).)).)))(((.(((((((((--(((.((.........((((........))))((((..((((......))))..))))-.((((....))-))..)).---)))))).)))))))))........------ ( -48.30, z-score = -1.89, R) >droSec1.super_11 1703780 157 - 2888827 GCUAUGAAAUUAUGGCACCGUGCCGCCAUAUUCGGUUGUCAGGGUUUCGUUAUUGAGU--GGCGACAUAAAUUAAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGGACCAUAAACUC-CUUGACAAGUC-AACAGUA---GCCACUCCAGUGAGAAGUUGGAAA------ ((((((....))))))(((.(((.(((......))).).)).)))(((.(((((((((--(((.((.........((((........))))((((..((((......))))..))))-.((((....))-))..)).---)))))).)))))))))........------ ( -48.30, z-score = -1.89, R) >droYak2.chr3L 5481381 156 + 24197627 GCUAUGAAAUUAUGGCACCGUGCUGCCAUAUUCGGUUGUCAGGGUUUCGUUAUUGAGU--GGCGACAUAAAUUAAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGAACCAUAAACUC-CUUGACAACUC-A-CAGUA---GCCACUCCACCGAGUAGUAGCAAA------ ((((((....))))))....((((((..((((((((......(((......)))((((--(((.((.........((((........))))((((..((((......))))..))))-...........-.-..)).---))))))).))))))))))))))..------ ( -49.60, z-score = -2.53, R) >droEre2.scaffold_4784 21458099 157 - 25762168 GCUAUGGAAUUAUGGCACCGUGCUGCCAUAUUCGGUGGUCAGGGUUUCGUUAUUGAGU--GGCGACAUAAAUUAAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGGACCAUAAACUC-CUUGACAAGUC-AGCAGUA---GCCACUCCACCGAGUAGUAGGAAA------ ((((((....)))))).((.((((((.....(((((((((((..........))))((--(((.((.........((((........))))((((..((((......))))..))))-.((((....))-))..)).---))))).)))))))))))))))...------ ( -50.60, z-score = -1.77, R) >droAna3.scaffold_13337 4098329 156 - 23293914 GCUAUGAAAUUAUGGCACCGUGUCGCCAUAUUCGGUGGUCGCUGUUUUGUUAUUGAGU--GGCGACAUAAAUUGAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGGACCAUAAACUC-CUUGACAAGUCUGCCAGCA--GGCUACUGUGCCUGCUCCGAAA--------- ..........((((((........))))))(((((..(((((..(((.......))).--.))))).........((((.(((((.(((..((((..((((......))))..))))-..))).)))))))))((((--(((......)))))))))))).--------- ( -56.80, z-score = -3.16, R) >dp4.chrXR_group6 8893687 160 + 13314419 GCUAUGAAAUUAUGGCACCGUAUCGCCAUAUUCGGUUGUUGGGGUUUCGUUAUUGAGU--GGCGACAUAAAUUAAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGGACCAUAAACUC-UAUGACAAGUG-AUCAGCU---GCAGUGGCAACUGGAAAAGGUUAAAAA--- ((((((....))))))(((((.(((((((..(((((...(((....)))..)))))))--)))))((((......((((........))))((((..((((......))))..))))-)))))).....-.((((.(---((....))).))))....)))......--- ( -43.70, z-score = -1.00, R) >droPer1.super_9 921861 160 + 3637205 GCUAUGAAAUUAUGGCACCGUAUCGCCAUAUUCGGUUGUUGGGGUUUCGUUAUUGAGU--GGCGACAUAAAUUAAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGGACCAUAAACUC-UAUGACAAGUG-AUCAGCU---GCAGUGGCAACUGGAAAAGGUUAAAAA--- ((((((....))))))(((((.(((((((..(((((...(((....)))..)))))))--)))))((((......((((........))))((((..((((......))))..))))-)))))).....-.((((.(---((....))).))))....)))......--- ( -43.70, z-score = -1.00, R) >droWil1.scaffold_180698 5234726 170 - 11422946 GCUAUGAAAUUAUGGCAUCACAUCACCAUAUUCGGUUGCUAGUAUUUCGUUAUUGAGUGGGGCAACAUAAAUUAAAUCAUGAUUUAUUGCCGAGUAGGUGGAAUAGACCAUAAAAUCAUGUGACAAGUCCAACAACAAAAGCAACAACUACUACUACGACUACCUGAGAA (((((((..((((((.......(((((.((((((((((((.....((((....))))...))))))((((((((.....))))))))...)))))))))))......))))))..))))(.((....))).........)))............................ ( -34.62, z-score = -0.28, R) >consensus GCUAUGAAAUUAUGGCACCGUGCCGCCAUAUUCGGUUGUCAGGGUUUCGUUAUUGAGU__GGCGACAUAAAUUAAGGCAUGAUUUAUUGCCGAGUAGGUGGAAUGGACCAUAAACUC_CUUGACAAGUC_AACAGUA___GCCACUCCAACGAGAAGUAGGAAA______ .........((((((..((((.(((((.(((((.((((((.....((((....))))...)))))).........((((........)))))))))))))).)))).))))))......................................................... (-32.84 = -32.42 + -0.41)

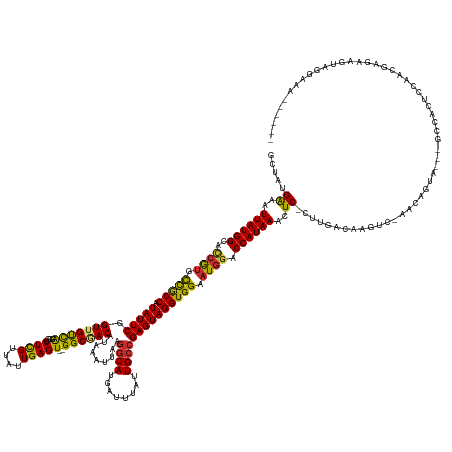

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:42:59 2011