
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 21,449,730 – 21,449,826 |
| Length | 96 |
| Max. P | 0.876810 |

| Location | 21,449,730 – 21,449,826 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 80.79 |
| Shannon entropy | 0.37125 |
| G+C content | 0.34900 |
| Mean single sequence MFE | -12.69 |
| Consensus MFE | -9.20 |
| Energy contribution | -9.20 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.876810 |
| Prediction | RNA |
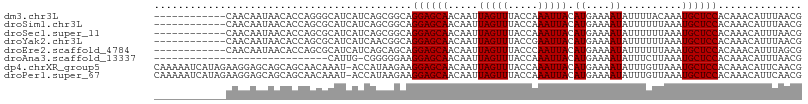
Download alignment: ClustalW | MAF
>dm3.chr3L 21449730 96 - 24543557 ------------CAACAAUAACACCAGGGCAUCAUCAGCGGCAGGAGCAACAAUUAGUUUACCAAAUUACAUGAAAAUAUUUUACAAAUGCUCCACAAACAUUUAACG ------------................((.......))....((((((.....(((((.....))))).((....))..........)))))).............. ( -9.90, z-score = -0.36, R) >droSim1.chr3L 20786180 96 - 22553184 ------------CAACAAUAACACCAGCGCAUCAUCAGCGGCAGGAGCAACAAUUAGUUUACCAAAUUACAUGAAAAUAUUUUUUAAAUGCUCCACAAACAUUUAACG ------------...............(((.......)))...((((((.....(((((.....)))))..(((((.....)))))..)))))).............. ( -12.60, z-score = -1.77, R) >droSec1.super_11 1373224 96 - 2888827 ------------CAACAAUAACACCAGCGCAUCAUCAGCGGCAGGAGCAACAAUUAGUUUACCAAAUUACAUGAAAAUAUUUUUUAAAUGCUCCACAAACAUUUAACG ------------...............(((.......)))...((((((.....(((((.....)))))..(((((.....)))))..)))))).............. ( -12.60, z-score = -1.77, R) >droYak2.chr3L 5126713 96 + 24197627 ------------CAACAAUAACACCAGCGCAUCAUCAACGGCAGGAGCAACAAUUAGUUUACCGAAUUACAUGAAAAUAUUUUUUAAAUGCUCCACAAACAUUUAACG ------------................((..........)).((((((.....((((((...))))))..(((((.....)))))..)))))).............. ( -10.40, z-score = -0.93, R) >droEre2.scaffold_4784 21107576 96 - 25762168 ------------CAACAAUAACACCAGCGCAUCAUCAGCAGCAGGAGCAACAAUUAGUUUACCCAAUUACAUGAAAAUAUUUUUUAAAUGCUCCACAAACAUUUAGCG ------------..............((((.......))....((((((.....(((((.....)))))..(((((.....)))))..))))))...........)). ( -13.10, z-score = -2.03, R) >droAna3.scaffold_13337 19891204 78 - 23293914 -----------------------------CAUUG-CGGGGGAAGGAGCAACAAUUAGUUUACCAAAUUACAUGAAAAUAUUUCUUAAAUGCUCCACAAACAUUUAACG -----------------------------.....-........((((((.....(((((.....)))))...(((.....))).....)))))).............. ( -9.50, z-score = -0.23, R) >dp4.chrXR_group5 22686 107 + 740492 CAAAAAUCAUAGAAGGAGCAGCAGCAACAAAU-ACCAUAAGAAGGAGCAACAAUUAGUUUACCAAAUUACAUGAAAAUAUUUGUUAAAUGCUCCACAAACAUUCAACG ..............((((((.....(((((((-(((.......)).....((..(((((.....)))))..))....))))))))...)))))).............. ( -16.70, z-score = -2.88, R) >droPer1.super_67 12174 107 - 321716 CAAAAAUCAUAGAAGGAGCAGCAGCAACAAAU-ACCAUAAGAAGGAGCAACAAUUAGUUUACCAAAUUACAUGAAAAUAUUUGUUAAAUGCUCCACAAACAUUCAACG ..............((((((.....(((((((-(((.......)).....((..(((((.....)))))..))....))))))))...)))))).............. ( -16.70, z-score = -2.88, R) >consensus ____________CAACAAUAACACCAGCGCAUCAUCAGCGGCAGGAGCAACAAUUAGUUUACCAAAUUACAUGAAAAUAUUUUUUAAAUGCUCCACAAACAUUUAACG ...........................................((((((.....(((((.....))))).((....))..........)))))).............. ( -9.20 = -9.20 + 0.00)

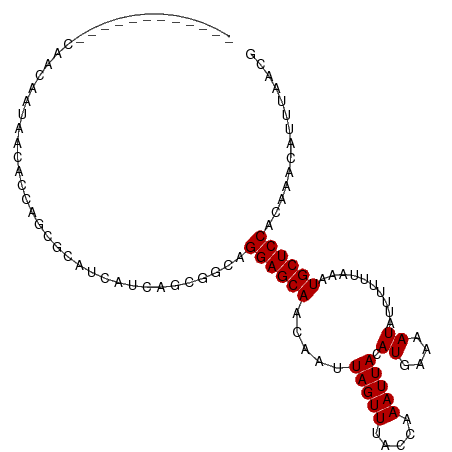
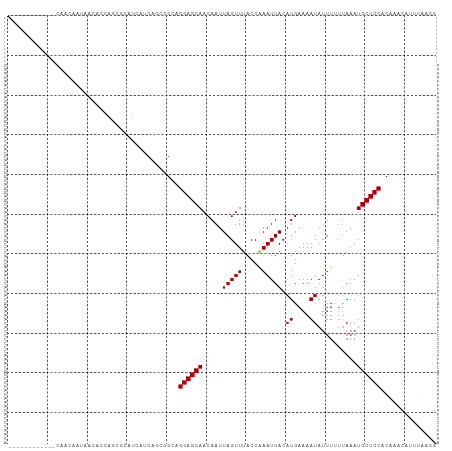
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:42:13 2011