
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 21,180,903 – 21,180,954 |
| Length | 51 |
| Max. P | 0.998875 |

| Location | 21,180,903 – 21,180,954 |
|---|---|
| Length | 51 |
| Sequences | 6 |
| Columns | 57 |
| Reading direction | forward |
| Mean pairwise identity | 77.36 |
| Shannon entropy | 0.40581 |
| G+C content | 0.35913 |
| Mean single sequence MFE | -12.75 |
| Consensus MFE | -9.14 |
| Energy contribution | -9.25 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.33 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.65 |
| SVM RNA-class probability | 0.993875 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
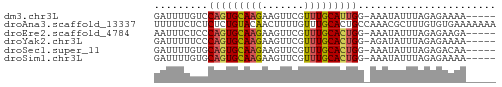
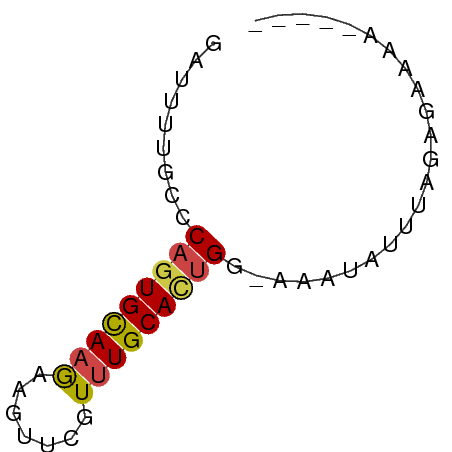
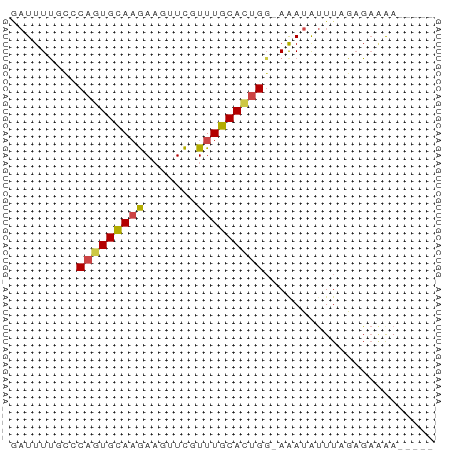
>dm3.chr3L 21180903 51 + 24543557 GAUUUUGUCCAGUGCAAGAAGUUCGUUUGCAUUGG-AAAUAUUUAGAGAAAA----- .......(((((((((((.......))))))))))-)...............----- ( -13.80, z-score = -3.42, R) >droAna3.scaffold_13337 14775430 57 - 23293914 UUUUUCUCUCUCUGUACAACUUUUGUUUGCACUGCCAAACGCUUUGUGUGAAAAAAA .....................(((.((..(((.((.....))...)))..)).))). ( -7.10, z-score = -0.69, R) >droEre2.scaffold_4784 20865534 51 + 25762168 AAUUUCUCCCAGUGCAAGAAGUUCGUUUGCACUGG-AAAUAUUUAGAGAAGA----- ..((((((((((((((((.......))))))))))-.........)))))).----- ( -17.40, z-score = -4.29, R) >droYak2.chr3L 4849866 51 - 24197627 GAUUUUUCCCAGUGCAAGAAGUUCGUUUGCACUGG-AGAUAUUUAGAGAAAA----- ......((((((((((((.......))))))))))-.)).............----- ( -15.40, z-score = -3.67, R) >droSec1.super_11 1134299 51 + 2888827 GAUUUUGUGCAGUGCAAGAAGUUCGUUUGCACUGG-AAAUAUUUAGAGACAA----- .........(((((((((.......))))))))).-................----- ( -11.40, z-score = -1.26, R) >droSim1.chr3L 20537331 51 + 22553184 GAUUUUGUGCAGUGCAAGAAGUUCGUUUGCACUGG-AAAUAUUUAGAGAAAA----- .........(((((((((.......))))))))).-................----- ( -11.40, z-score = -1.53, R) >consensus GAUUUUGCCCAGUGCAAGAAGUUCGUUUGCACUGG_AAAUAUUUAGAGAAAA_____ .........(((((((((.......)))))))))....................... ( -9.14 = -9.25 + 0.11)
| Location | 21,180,903 – 21,180,954 |
|---|---|
| Length | 51 |
| Sequences | 6 |
| Columns | 57 |
| Reading direction | reverse |
| Mean pairwise identity | 77.36 |
| Shannon entropy | 0.40581 |
| G+C content | 0.35913 |
| Mean single sequence MFE | -11.90 |
| Consensus MFE | -6.48 |
| Energy contribution | -7.32 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.66 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.53 |
| SVM RNA-class probability | 0.998875 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
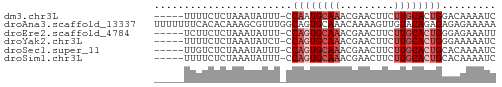
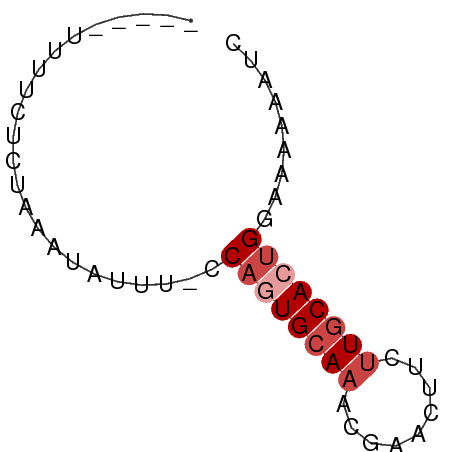
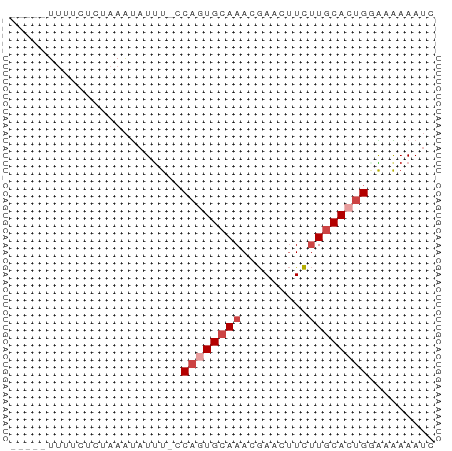
>dm3.chr3L 21180903 51 - 24543557 -----UUUUCUCUAAAUAUUU-CCAAUGCAAACGAACUUCUUGCACUGGACAAAAUC -----...............(-(((.(((((.........))))).))))....... ( -8.90, z-score = -3.89, R) >droAna3.scaffold_13337 14775430 57 + 23293914 UUUUUUUCACACAAAGCGUUUGGCAGUGCAAACAAAAGUUGUACAGAGAGAGAAAAA ((((((((.......((.....)).((((((.......)))))).)))))))).... ( -12.00, z-score = -1.54, R) >droEre2.scaffold_4784 20865534 51 - 25762168 -----UCUUCUCUAAAUAUUU-CCAGUGCAAACGAACUUCUUGCACUGGGAGAAAUU -----.............(((-(((((((((.........))))))))))))..... ( -16.30, z-score = -4.71, R) >droYak2.chr3L 4849866 51 + 24197627 -----UUUUCUCUAAAUAUCU-CCAGUGCAAACGAACUUCUUGCACUGGGAAAAAUC -----.............((.-(((((((((.........)))))))))))...... ( -14.80, z-score = -5.19, R) >droSec1.super_11 1134299 51 - 2888827 -----UUGUCUCUAAAUAUUU-CCAGUGCAAACGAACUUCUUGCACUGCACAAAAUC -----................-.((((((((.........))))))))......... ( -9.70, z-score = -2.98, R) >droSim1.chr3L 20537331 51 - 22553184 -----UUUUCUCUAAAUAUUU-CCAGUGCAAACGAACUUCUUGCACUGCACAAAAUC -----................-.((((((((.........))))))))......... ( -9.70, z-score = -3.66, R) >consensus _____UUUUCUCUAAAUAUUU_CCAGUGCAAACGAACUUCUUGCACUGGAAAAAAUC .......................((((((((.........))))))))......... ( -6.48 = -7.32 + 0.83)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:41:45 2011