
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 21,041,708 – 21,041,904 |
| Length | 196 |
| Max. P | 0.864171 |

| Location | 21,041,708 – 21,041,799 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 73.59 |
| Shannon entropy | 0.46017 |
| G+C content | 0.43217 |
| Mean single sequence MFE | -20.60 |
| Consensus MFE | -9.10 |
| Energy contribution | -10.13 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr3L 21041708 91 + 24543557 --AUUUAUUUAACUA-AUACAUCACUGAAGCGAAUCAACUAACUAGGGACUUAGUUAGUGGGACCAGCUUUAGUCCCCCCA--AUUCGCCACCCCC--------- --.............-.............((((((..(((((((((....)))))))))(((((........)))))....--)))))).......--------- ( -22.60, z-score = -3.02, R) >droSim1.chr3L 20395796 91 + 22553184 --AUUUAUUUACCUA-AUACAUCACUGAAGCGAAUCGACUAACUAGGGACUUAGUUAGUGGGACCUGCUUUAGUCCACCCA--AUUCGCCACCCCC--------- --.............-.......(((((((((..((.(((((((((....))))))))).))...))))))))).......--.............--------- ( -21.30, z-score = -2.23, R) >droSec1.super_11 930230 91 + 2888827 --AUUUAUUUACCUA-AUACAUCACUGAAGCGAAUCGACUAACUAGGGACUUAGUUAGUGGGACCUGCUUUACUCCACCCA--AUUCGCCACCCCC--------- --.............-.............((((((..(((((((((....)))))))))(((...............))).--)))))).......--------- ( -17.76, z-score = -1.48, R) >droYak2.chr3L 4711161 91 - 24197627 --AUUUAUUUACCUA-AUACAUAAUUGAAGAGGAGCAACUAACUAGGGAAUUAGUUAGUGGGAGCUGCUUAAGUUCCCCCA--AUUCGCCACCCCC--------- --.............-..........(((..((....(((((((((....)))))))))(((((((.....))))))))).--.))).........--------- ( -23.60, z-score = -1.97, R) >droEre2.scaffold_4784 20723656 92 + 25762168 --AUUUAUUUACCUA-ACACCUAAUUGAAGCGGAUCAACUAACUAGGGACUUAGUUAGUGGGACCUCUUUUAGUCCCCUCA--AUUCGCCACCCCCA-------- --.............-.............((((((..(((((((((....)))))))))(((((........)))))....--))))))........-------- ( -22.10, z-score = -2.03, R) >droWil1.scaffold_180949 1023991 105 - 6375548 CUAACUAUUUACCCGUAUACAUAAUUGAAGAGGAGGAUAGUGCAGGAGGAAGAGGAAUUAGGAAAAGGGAUAGAGAAAGCAGGAUAUCCUAUAUCCAGAACCAGA ((.((((((..(((...............).))..))))))..))........((.....(((.(.((((((............)))))).).)))....))... ( -16.26, z-score = 0.30, R) >consensus __AUUUAUUUACCUA_AUACAUAACUGAAGCGAAUCAACUAACUAGGGACUUAGUUAGUGGGACCUGCUUUAGUCCACCCA__AUUCGCCACCCCC_________ .............................(((((...(((((((((....)))))))))(((...............)))....)))))................ ( -9.10 = -10.13 + 1.03)



| Location | 21,041,799 – 21,041,904 |
|---|---|
| Length | 105 |
| Sequences | 10 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 82.10 |
| Shannon entropy | 0.38165 |
| G+C content | 0.42876 |
| Mean single sequence MFE | -29.59 |
| Consensus MFE | -22.87 |
| Energy contribution | -22.79 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.864171 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 21041799 105 - 24543557 AAGCCACUUACCAGUAAAUCUCGUUGGAGGUAAUCAUAUUGGUGGUCUUGUAAUACUGGGCGGCAGUGUAGCUGCUCAUUCUGCCG-UGUGUUUUUGUUGUUUUCU--- ....(((...((((((.(((((....))))).(((((....))))).......))))))(((((((.((........)).))))))-))))...............--- ( -27.20, z-score = -0.81, R) >droSim1.chr3L 20395887 103 - 22553184 AAGCCACUUACCAGUAAAUCUCAUUGGAGGUAAUCAUAUUGGUGGUCUUGUAAUACUGGGCGGCAGUGUAGCUGCUCAUUCUGCCG--ACG-UGUUGUUGUUUUCU--- ....(((.((((((((.(((((....))))).....))))))))(((..(((....((((((((......))))))))...))).)--)))-))............--- ( -27.90, z-score = -0.73, R) >droSec1.super_11 930321 103 - 2888827 AAGCCACUUACCAGUAAAUCUCAUUGGAGGUAAUCAUAUUGGUGGUCUUGUAAUACUGGGCGGCAGUGUAGCUGCUCAUUCUGCCG--ACG-UGUUGUUGUUUUCU--- ....(((.((((((((.(((((....))))).....))))))))(((..(((....((((((((......))))))))...))).)--)))-))............--- ( -27.90, z-score = -0.73, R) >droYak2.chr3L 4711252 103 + 24197627 GGGCAACUUACCAGUAAAUCUCAUUGGAGGUAAUCAUAUUGGUGGUCUUGUAAUACUGGGCGGCAGUGUAGCUGCUCAUUCUGCCG--AUG-UGUUGUUGUUUUCU--- (((((((..........(((((....)))))...((((((((..(...........((((((((......))))))))..)..)))--)))-))..)))))))...--- ( -31.62, z-score = -1.87, R) >droEre2.scaffold_4784 20723748 106 - 25762168 AAGCAACUUACCAGUAAAUCUCAUUGGAGGUAAUCAUAUUGGUGGUCUUGUAAUACUGGGCGGCAGUGUAGCUGCUCAUUCUGCCG--AUG-UGUUGUUGUUUUUUUUG (((((((..........(((((....)))))...((((((((..(...........((((((((......))))))))..)..)))--)))-))..)))))))...... ( -32.12, z-score = -2.51, R) >droAna3.scaffold_13337 14642331 94 + 23293914 -----ACUUACCAGUAUAUCUCAUUGGAGGUAAUCAUAUUGGUGGUCUUGUAGUACUGGGCGGCAGUAUAGCUGCUCAUUCUGACG--GUCUAUGUUCUAU-------- -----....(((((((((((((....))))))....)))))))(..(...(((...((((((((......))))))))..)))..)--..)..........-------- ( -27.50, z-score = -1.61, R) >dp4.chrXR_group6 7346632 105 - 13314419 AGGACACUUACCAGUAGAUCUCAUUGGAGGUAAUCAUAUUGGUGGUCUUGUAGUACUGGGCGGCAGUGUAGCUGCUCAUUGUGUUU-CA-GCUGCUGUCAUUUACUU-- (((((...((((((((.(((((....))))).....)))))))))))))..((((...(((((((((..(((.((.....))))).-..-)))))))))...)))).-- ( -34.80, z-score = -1.91, R) >droPer1.super_41 702986 105 - 728894 AGGACACUUACCAGUAGAUCUCAUUGGAGGUAAUCAUAUUGGUGGUCUUGUAGUACUGGGCGGCAGUGUAGCUGCUCAUUGUGUUU-CA-GCUGCUGUCAUUUACUU-- (((((...((((((((.(((((....))))).....)))))))))))))..((((...(((((((((..(((.((.....))))).-..-)))))))))...)))).-- ( -34.80, z-score = -1.91, R) >droWil1.scaffold_180949 1024096 104 + 6375548 -----ACUUACCAGUAGAUUUCAUUAGAGGUAAUCAUAUUGGUGGUCUUGUAGUACUGAGCGGCCGUAUAGCUGCUCAUUCUGCUUCAAUAGAAACGUUGUGCUUUUGU -----..(((((((((((((((......)).)))).)))))))))......(((((((((((((......))))))))(((((......))))).....)))))..... ( -29.00, z-score = -1.66, R) >droVir3.scaffold_13049 9935927 100 + 25233164 -----ACUUACCAGUAAAUCUCAUUCGAGGUAAUCAUAUUGGUGGUCUUGUAGUACUGGGCUGCCGUGUAGCUGCUCAUUGUGUAG--AUAUAUAUAGUAUAUAAAU-- -----..(((((((((.(((((....))))).....)))))))))..(((((.((((((((.((......)).))))).((((((.--...))))))))))))))..-- ( -23.10, z-score = -0.30, R) >consensus AAGCCACUUACCAGUAAAUCUCAUUGGAGGUAAUCAUAUUGGUGGUCUUGUAAUACUGGGCGGCAGUGUAGCUGCUCAUUCUGCCG__AUG_UGUUGUUGUUUUCU___ .......(((((((((.(((((....))))).....)))))))))...........((((((((......))))))))............................... (-22.87 = -22.79 + -0.08)


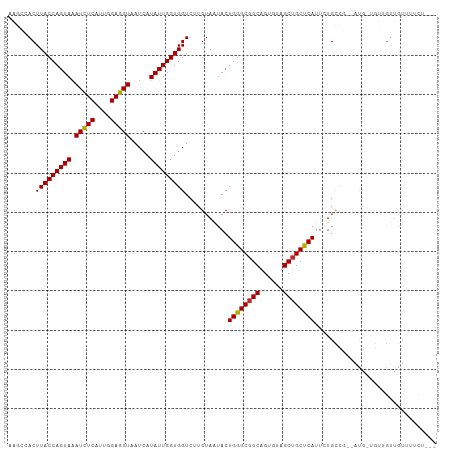
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:41:22 2011