
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 21,004,118 – 21,004,227 |
| Length | 109 |
| Max. P | 0.735450 |

| Location | 21,004,118 – 21,004,227 |
|---|---|
| Length | 109 |
| Sequences | 8 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 68.03 |
| Shannon entropy | 0.63036 |
| G+C content | 0.45607 |
| Mean single sequence MFE | -21.06 |
| Consensus MFE | -6.54 |
| Energy contribution | -6.60 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.735450 |
| Prediction | RNA |
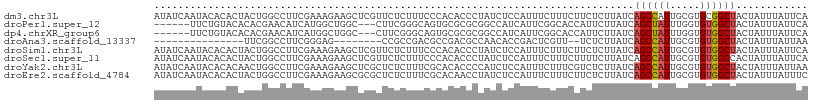
Download alignment: ClustalW | MAF
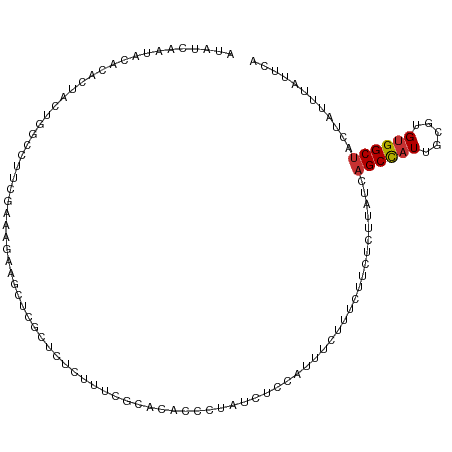
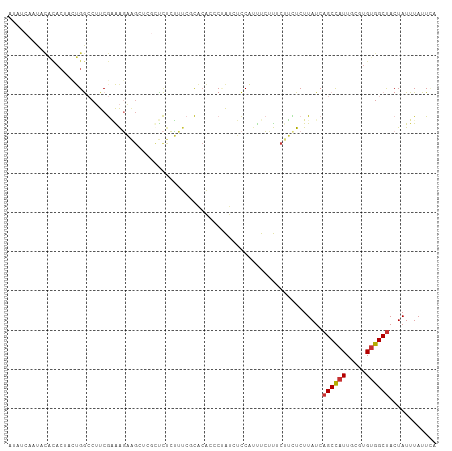
>dm3.chr3L 21004118 109 - 24543557 AUAUCAAUACACACUACUGGCCUUCGAAAGAAGCUCGUUCUCUUUCCCACACCCUAUCUCCAUUUCUUUCUUCUCUUAUCAGCCAUUGCGUGCGGCUACUAUUUAUUCA .................((((((((....)))((.(((.................................................))).)))))))........... ( -13.19, z-score = -1.09, R) >droPer1.super_12 590477 100 + 2414086 ------UUCUGUACACACGAACAUCAUGGCUGGC---CUUCGGGCAGUGCGCGCGGCCAUCAUUCGGCACCAUUCUUAUCAGCUAUUUGGUGUGGCUACUAUUUAUUCA ------.......(((((.((....(((((((((---(....))).......(..(((.......)))..)........))))))))).)))))............... ( -28.80, z-score = -1.32, R) >dp4.chrXR_group6 12178918 100 + 13314419 ------UUCUGUACACACGAACAUCAUGGCUGGC---CUUCGGGCAGUGCGCGCGGCCAUCAUUCGGCACCAUUCUUAUCAGCUAUUUGGUGUGGCUACUAUUUAUUCA ------.......(((((.((....(((((((((---(....))).......(..(((.......)))..)........))))))))).)))))............... ( -28.80, z-score = -1.32, R) >droAna3.scaffold_13337 14611537 84 + 23293914 ---------------UUCGGCCUUCGGGAG--------CCGCCGACGCCGACGCCAACACCGACUCGUU--UCUCUUAUCAGCCAUUGCGUGUGGCUACUAUUUAUUAA ---------------.(((((..((((...--------...)))).))))).((((.((((((...(((--.........)))..))).)))))))............. ( -21.80, z-score = -0.38, R) >droSim1.chr3L 20359158 109 - 22553184 AUAUCAAUACACACUACUGGCCUUCGAAAGAAGCUCGUUCUCUUUCCCACACCCUAUCUCCAUUUCUUUCUUCUCUUAUCAGCCAUUGCGUGUGGCUACUAUUUAUUCA .........(((((((.((((((((....))))...((..........))...............................)))).)).)))))............... ( -16.70, z-score = -2.52, R) >droSec1.super_11 900925 109 - 2888827 AUAUCAAUACACACUACUGGCCUUCGAAAGAAGCUCGUUCUCUUUCCCACACCCUAUCUCCAUUUCUUUCUUUUCUUAUCAGCCAUUGCGUGUGGCCACUAUUUAUUCA .........(((((((.((((((((....))))...((..........))...............................)))).)).)))))............... ( -16.70, z-score = -2.51, R) >droYak2.chr3L 4679217 109 + 24197627 AUAUCAAUACACACAACUGGCCUUCGAAAGAAGCUCGCUCUCUUUCGCACACCCCAUCUCCAUUUCUUUCGUCUCUUAUCAGCCAUUGCGUGUGGCUACUAUUUAUUAA .......(((((.(((.((((((((....))))...((........)).................................))))))).)))))............... ( -20.90, z-score = -3.04, R) >droEre2.scaffold_4784 20693609 109 - 25762168 AUAUCAAUACACACUACUGGCCUUCGAAAGAAGCGCGCUCUCUUUCGCACAACCUAUCUCCAUUUCUUUCUUCUCUUAUCAGCCAUUGCGUGUGGCUACUAUUUAUUUC .........(((((((.((((((((....)))).(((........))).................................)))).)).)))))............... ( -21.60, z-score = -3.24, R) >consensus AUAUCAAUACACACUACUGGCCUUCGAAAGAAGCUCGCUCUCUUUCGCACACCCUAUCUCCAUUUCUUUCUUCUCUUAUCAGCCAUUGCGUGUGGCUACUAUUUAUUCA ................................................................................((((((.....))))))............ ( -6.54 = -6.60 + 0.06)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:41:15 2011