
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 20,977,418 – 20,977,514 |
| Length | 96 |
| Max. P | 0.724130 |

| Location | 20,977,418 – 20,977,514 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 72.59 |
| Shannon entropy | 0.55189 |
| G+C content | 0.60106 |
| Mean single sequence MFE | -29.90 |
| Consensus MFE | -16.86 |
| Energy contribution | -16.48 |
| Covariance contribution | -0.39 |
| Combinations/Pair | 1.57 |
| Mean z-score | -1.00 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.724130 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
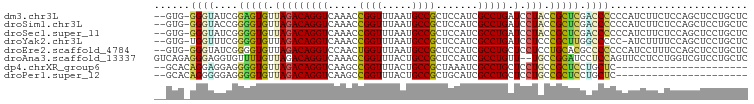
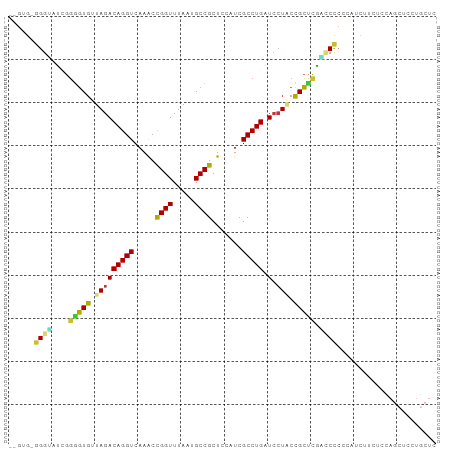
>dm3.chr3L 20977418 96 + 24543557 --GUG-GGGUAUCGGAGUGUUAGACAGGUCAAACCGGUUUAAUGCCGCUCCAUCGCCUGAUCCUACCGCUCGACCCCCCAUCUUCUCCAGCUCCUGCUC --(((-(((..(((.((((.(((.(((((.....((((.....)))).......)))))...))).)))))))..)))))).......(((....))). ( -32.20, z-score = -2.51, R) >droSim1.chr3L 20331995 96 + 22553184 --GUG-GGGUACCGGGGUGUUAGACAGGUCAAACCGGUUUAAUGCCGCUCCAUCGCCUGAUCCUACCGCUCGACCCCCCAUCUUCUCCAGCUCCUGCUC --(.(-((((..(((((((((((((.((.....)).))))))))))((......)).........)))....))))).).........(((....))). ( -30.00, z-score = -0.87, R) >droSec1.super_11 873821 96 + 2888827 --GUG-GGGUAUCGGGGUGUUAGACAGGUCAAACCGGUUUAAUGCCGCUCCAUCGCCUGAUCCUACCGCUCGACCCCCCAUCUUCUCCAGCUCCUGCUC --(((-(((..((((((.((..(((((((.....((((.....)))).......))))).))..))).)))))..)))))).......(((....))). ( -32.50, z-score = -1.84, R) >droYak2.chr3L 4640198 95 - 24197627 --GUG-UGGUUUCGGGGUGUUAGACAGGUCAAACCGGUUUAAUGCCGCUCCAUCGCCUGAUCCUCCCGCUUGGCCCCC-AUCUUUUCCAGCUCCUGCUC --(((-.((..((((((((((((((.((.....)).))))))))))((......))))))....))))).........-.........(((....))). ( -24.90, z-score = 0.06, R) >droEre2.scaffold_4784 20666249 96 + 25762168 --GUG-GGGUAUCGGGGUGUUAGACAGGUCCAACUGGUUUAAUGCCGCUCCAUCGCCUGCUCCUCCUGCACGCCCCCCCAUCCUUUCCAGCUCCUGCUC --(((-(((....(.((((((((((((......)).)))))))))).)......((.(((.......))).))..)))))).......(((....))). ( -29.90, z-score = -0.78, R) >droAna3.scaffold_13337 14582562 97 - 23293914 GUCAGAGGGAGGUGUUUUGUUAGACAGGUCAAACCGGUUUACUGCCGCUCCAUCGCCUGUU--UGCCGGAUCCUCCAGUUCCUCCUGGUCGUCCUGCUC (.(((..(((((...((((.(((((((((.....((((.....)))).......)))))))--)).)))).)))))....((....)).....)))).. ( -29.20, z-score = -0.49, R) >dp4.chrXR_group6 12151097 75 - 13314419 --GCACAGGAGGAGGGGUGUUAGACAGGUCAAGCCGGUUUACUGCCGCUAAAUCGCCUGCUCCUGCCGCUCCUGCUC---------------------- --....((.(((((.((((..((.(((((..(((.(((.....)))))).....)))))))..)))).))))).)).---------------------- ( -29.60, z-score = -1.13, R) >droPer1.super_12 562125 75 - 2414086 --GCACAGGGGGAGGGGUGUUAGACAGGUCAAGCCGGUUUACUGCCGCUGCAUCGCCUGCUCCUGCCGCUCCUGCUC---------------------- --....((.(((((.((((..((.(((((...((((((.....))))..))...)))))))..)))).))))).)).---------------------- ( -30.90, z-score = -0.42, R) >consensus __GUG_GGGUAUCGGGGUGUUAGACAGGUCAAACCGGUUUAAUGCCGCUCCAUCGCCUGAUCCUACCGCUCGACCCCCCAUCUUCUCCAGCUCCUGCUC ......((((....(((((.(((.(((((.....((((.....)))).......)))))...))).))))).))))....................... (-16.86 = -16.48 + -0.39)
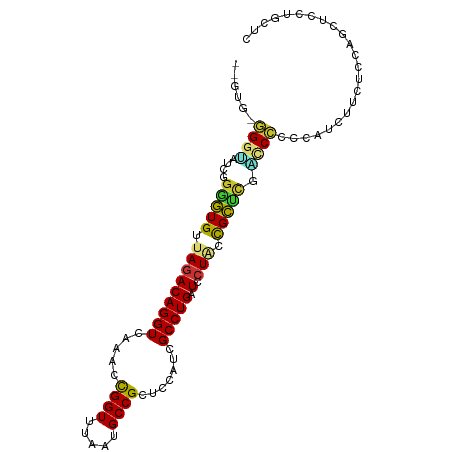

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:41:11 2011