| Sequence ID | dm3.chr3L |
|---|---|
| Location | 20,901,244 – 20,901,346 |
| Length | 102 |
| Max. P | 0.957234 |

| Location | 20,901,244 – 20,901,346 |
|---|---|
| Length | 102 |
| Sequences | 8 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 75.85 |
| Shannon entropy | 0.45014 |
| G+C content | 0.49737 |
| Mean single sequence MFE | -29.20 |
| Consensus MFE | -19.61 |
| Energy contribution | -19.12 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.957234 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
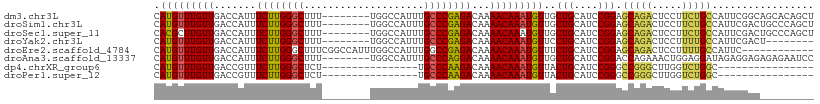
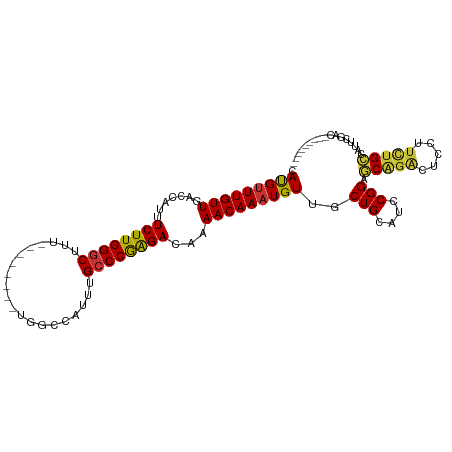
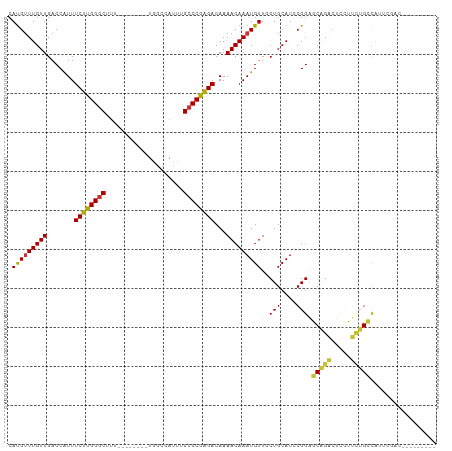
>dm3.chr3L 20901244 102 + 24543557 CAUGUUUGUUGACCAUUUCUUGGGCUUU--------UGGCCAUUUGCCCGAGACAAAACAAAUGUUGCUGCAUCCGGAGCAGACUCCUUCUGCCAUUCGGCAGCACAGCU ...(((((((.......((((((((...--------.........))))))))...)))))))((((.(((..((((((((((.....)))))..)))))..))))))). ( -34.80, z-score = -2.24, R) >droSim1.chr3L 20260347 102 + 22553184 CAUGUUUGUUGACCAUUUCUUGGGCUUU--------UGGCCAUUUGCCCGAGACAAAACAAAUGUUGCUGCAUCCGGAGCAGACUCCUUCUGCCAUUCGACUGCCCAGCU ...(((((((.......((((((((...--------.........))))))))...)))))))((((..(((..(((((((((.....)))))..))))..))).)))). ( -31.20, z-score = -2.00, R) >droSec1.super_11 797777 102 + 2888827 CACGCUUGUUGACCAUUUCUUGGGCUUU--------UGGCCAUUUGCCCGAGACAAAACAAAUGUUGCUGCAUCCGGAGCAGACUCCUUCUGCCAUUCGACUGCCCAGCU .(((.(((((.......((((((((...--------.........))))))))...))))).))).((((((..(((((((((.....)))))..))))..)))..))). ( -29.50, z-score = -1.41, R) >droYak2.chr3L 4558769 94 - 24197627 CAUGUUUGUUGACCAUUUCUUGGGCUUU--------UGGCCAUUUGCCCGAGACAAAACAAAUGUUCCUGCAUCCGGAGCAGACUCCUUUUGCCAUUCGACU-------- .......(((((.....((((((((...--------.........))))))))(((((....((((((.......))))))......)))))....))))).-------- ( -25.10, z-score = -1.63, R) >droEre2.scaffold_4784 20589509 98 + 25762168 CAUGUUUGUUGACCAUUUCUUGGGCUUUCGGCCAUUUGGCCAUUUGGCCGAGACAAAACAAAUGUUUCUGCAUCCGGAGCAGACUCCUUUUGCCAUUC------------ .(((((((((..(((.....)))..(((((((((..(....)..)))))))))...)))))))))....(((...((((....))))...))).....------------ ( -29.50, z-score = -1.89, R) >droAna3.scaffold_13337 14509365 102 - 23293914 CAUGUUUGUUGACCAUUUCUUGGGCUUU--------UGGCCAUUUGCCCAGGACAAAACAAAUGUUGCUGCAUCCGGACCAGAAACUGGAGGAUAGAGGAGAGAGAAUCC .(((((((((.......((((((((...--------.........))))))))...)))))))))......((((...((((...)))).))))...(((.......))) ( -25.70, z-score = 0.10, R) >dp4.chrXR_group6 12077947 78 - 13314419 CAUGUUUGUUGACCGUUUCUUGGGCUCU----------------UGCCCAAGACAAAACAAAUGUUACUGCAUCCGGGCCGGGCUUGGUCUGGC---------------- .(((((((((.......((((((((...----------------.))))))))...)))))))))........((((((((....)))))))).---------------- ( -28.90, z-score = -2.41, R) >droPer1.super_12 489017 78 - 2414086 CAUGUUUGUUGACCGUUUCUUGGGCUCU----------------UGCCCAAGACAAAACAAAUGUUACUGCAUCCGGGCCGGGCUUGGUCUGGC---------------- .(((((((((.......((((((((...----------------.))))))))...)))))))))........((((((((....)))))))).---------------- ( -28.90, z-score = -2.41, R) >consensus CAUGUUUGUUGACCAUUUCUUGGGCUUU________UGGCCAUUUGCCCGAGACAAAACAAAUGUUGCUGCAUCCGGAGCAGACUCCUUCUGCCAUUCGAC_________ .(((((((((.......((((((((....................))))))))...)))))))))..(((....))).(((((.....)))))................. (-19.61 = -19.12 + -0.48)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:41:04 2011