
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 20,689,633 – 20,689,727 |
| Length | 94 |
| Max. P | 0.577108 |

| Location | 20,689,633 – 20,689,727 |
|---|---|
| Length | 94 |
| Sequences | 8 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 77.36 |
| Shannon entropy | 0.42687 |
| G+C content | 0.32092 |
| Mean single sequence MFE | -18.95 |
| Consensus MFE | -10.01 |
| Energy contribution | -9.71 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.577108 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 20689633 94 - 24543557 CAAUAAAUAUUCAUAAAAAGAGUUAAGUGCCGCCAUAAGGCUCUUUGUUUAC--ACAAAGGAGAAAUUAUGUUGGAAGUUGACUUUUA--AAAGGGUU .................(((((((((.(.(((.(((((..((((((((....--))))).)))...))))).))).).))))))))).--........ ( -21.70, z-score = -2.40, R) >droEre2.scaffold_4784 20382272 94 - 25762168 CAAUAAAUAUUCAUAAAAAGAGUUAAGUGCCGUCAUAAGGUUCUUUGUUUGG--ACACAGGAGAAAUUGUGUUGGAAGUUGACUUUUA--AAAGGGUA .......(((((.....(((((((((.(.(((.(((((..((((((((....--..))).))))).))))).))).).))))))))).--...))))) ( -20.10, z-score = -1.82, R) >droYak2.chr3L 4338280 94 + 24197627 CAAUAAAUAUUCAUAAAAAGAGUUAAGUGCCGUCAUAAGGCUCUUUGUUAGG--ACAAAGGAGAAAUUUUGUUGGAAGUUGACUUUUA--AAAGGGUU .................(((((((((.(.(((.((.((..((((((((....--))))).)))...)).)).))).).))))))))).--........ ( -18.20, z-score = -1.28, R) >droSec1.super_11 579000 94 - 2888827 CUAUAAAUAUUCGUAAAAAGAGUUAAGUGCCGCCAUAAGGCUCUUUGUUUAC--ACAAAGGAGAAAUUAUGUUGGAAGUUGACUUUUA--AAAGGGUU ........((((.....(((((((((.(.(((.(((((..((((((((....--))))).)))...))))).))).).))))))))).--...)))). ( -21.90, z-score = -2.15, R) >droSim1.chr3L 20043645 94 - 22553184 CAAUAAAUAUUCGUAAAAAGAGUUAAGUGCCGCCAUAAGGCUCUUUGUUUAC--ACAAAGGAGAAAUUAUGUUGGAAGUUGACUUUUA--AAAGGGUU ........((((.....(((((((((.(.(((.(((((..((((((((....--))))).)))...))))).))).).))))))))).--...)))). ( -21.90, z-score = -2.36, R) >dp4.chrXR_group8 8114776 86 - 9212921 ----------UCAUAAAAAGAGUUAAGUGCUACCAUAAGGUUCUUUUGUAUGCAGAAAAUAGGAAAUGAAGCUGAAAAUUGACUUUAG--UUGUGGCC ----------((((((..((((((((..((((((....)))((((((.(((.......))).)))).)))))......))))))))..--)))))).. ( -16.80, z-score = -1.09, R) >droPer1.super_40 306171 86 - 810552 ----------UCAUAAAAAGAGUUAAGUGCUACCAUAAGGUUCUUUUGUAUGCAGAAAAUAGGAAAUGAAGCUGAAAAUUGACUUUAG--UUGUGGCC ----------((((((..((((((((..((((((....)))((((((.(((.......))).)))).)))))......))))))))..--)))))).. ( -16.80, z-score = -1.09, R) >droAna3.scaffold_13337 11041516 98 - 23293914 CAAUAAAUAUUCAUAAAAAGAGUUAAGUGCUACCAUAAGAUUCUUUGUUUAGGAAAAAAGGGGUAAUAAUAUUUGUAGUUGACUUUAAAGAAAGGGUU ..................((((((((.(((((((......(((((.....)))))......)))).........))).))))))))............ ( -14.20, z-score = -0.54, R) >consensus CAAUAAAUAUUCAUAAAAAGAGUUAAGUGCCGCCAUAAGGCUCUUUGUUUAG__ACAAAGGAGAAAUUAUGUUGGAAGUUGACUUUUA__AAAGGGUU ..................((((((((.(.(((((....)))((((((........))))))............)).).))))))))............ (-10.01 = -9.71 + -0.30)
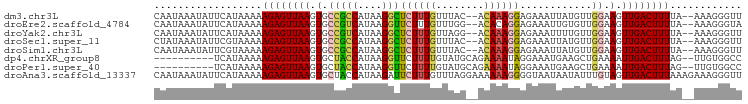
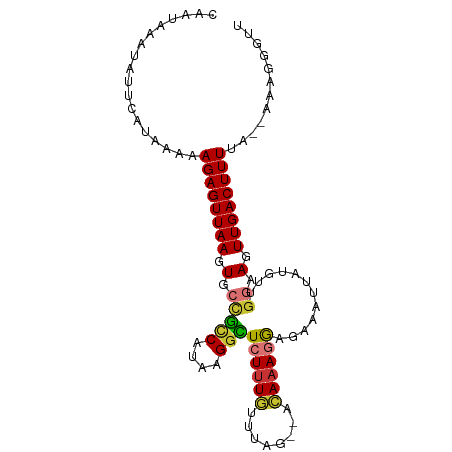

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:40:33 2011