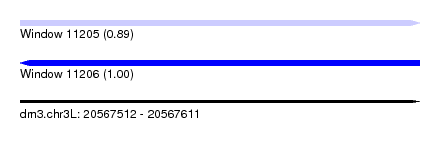

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 20,567,512 – 20,567,611 |
| Length | 99 |
| Max. P | 0.998394 |

| Location | 20,567,512 – 20,567,611 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 72.40 |
| Shannon entropy | 0.51027 |
| G+C content | 0.53410 |
| Mean single sequence MFE | -28.88 |
| Consensus MFE | -10.16 |
| Energy contribution | -10.88 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.51 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.892634 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
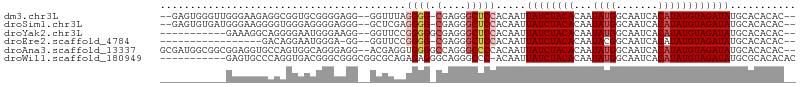
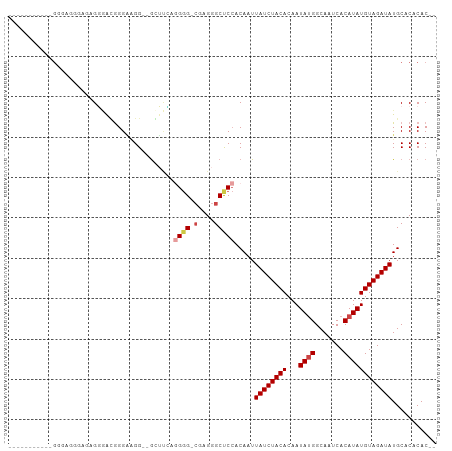
>dm3.chr3L 20567512 99 + 24543557 --GAGUGGGUUGGGAAGAGGCGGUGCGGGGAGG--GGUUUAGGGG-CGAGGGCUCCACAAUUAUCUACACAAUAUGGCAAUCACAUAUGUAGAUAUGCACACAC-- --..(((.(((.......))).(((((......--......((((-(....))))).....((((((((...((((.......))))))))))))))))).)))-- ( -25.90, z-score = -1.95, R) >droSim1.chr3L 19919848 99 + 22553184 --GAGUGUGAUGGGAAGGGGUGGGAGGGGAGGG--GCUCGAGGGG-CGAGGGCUCCACAAUUAUCUACACAAUAUGGCAAUCACAUAUGUAGAUAUGCACACAC-- --..(((((.....................(((--((((......-...))))))).....((((((((...((((.......))))))))))))..)))))..-- ( -28.40, z-score = -2.45, R) >droYak2.chr3L 4212390 91 - 24197627 -----------GAAAGGCAGGGGAAUGGGAAGG--GGUUCCGGGGGCGAGGGCUCCACAAUUAUCUACACAAUAUGGCAAUCACAUAUGUAGAUAUGCACACAC-- -----------.....(((..(((((.......--.))))).(((((....))))).....((((((((...((((.......)))))))))))))))......-- ( -24.80, z-score = -2.87, R) >droEre2.scaffold_4784 20261932 83 + 25762168 -----------------GACAGGAAUGGGA-GG--GGUUCCGGGG-CGAGGGCUCCACAAUUAUCUACACAAUACGGCAAUCACAUAUGUAGAUAUGCACACAC-- -----------------....(((((....-..--.)))))((((-(....))))).....((((((((..................)))))))).........-- ( -18.37, z-score = -1.24, R) >droAna3.scaffold_13337 10920363 102 + 23293914 GCGAUGGCGGCGGAGGUGCCAGUGGCAGGGAGG--ACGAGGUGGGGCCAGGGCCCCACAAUUAUCUACACAAUAUGGCAAUCACAUAUGUAGAUAUGCACACAC-- ....(((((.(...).)))))((((((......--.....(((((((....)))))))...((((((((...((((.......))))))))))))))).)))..-- ( -39.50, z-score = -2.98, R) >droWil1.scaffold_180949 4130810 94 - 6375548 -----------GAGUGCCCAGGUGACGGGCGGGCGGCGCAGAGAGGGCAGGGCCC-ACAAUUAUCUACACAAUAUGGCAAUCACAUAUGUAGAUAUGCGCACACAC -----------(..(((((.(....))))))..).(((((....((((...))))-.....((((((((...((((.......)))))))))))))))))...... ( -36.30, z-score = -3.60, R) >consensus ___________GGGAGGGAGAGGGACGGGAAGG__GCUUCAGGGG_CGAGGGCUCCACAAUUAUCUACACAAUAUGGCAAUCACAUAUGUAGAUAUGCACACAC__ .........................................((((.(....))))).....((((((((...((((.......))))))))))))........... (-10.16 = -10.88 + 0.72)
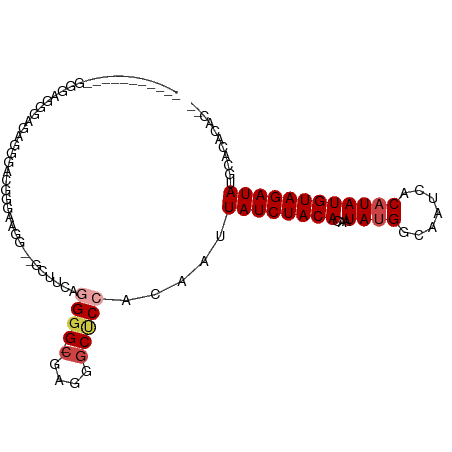

| Location | 20,567,512 – 20,567,611 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 72.40 |
| Shannon entropy | 0.51027 |
| G+C content | 0.53410 |
| Mean single sequence MFE | -27.80 |
| Consensus MFE | -13.42 |
| Energy contribution | -13.28 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.16 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.34 |
| SVM RNA-class probability | 0.998394 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
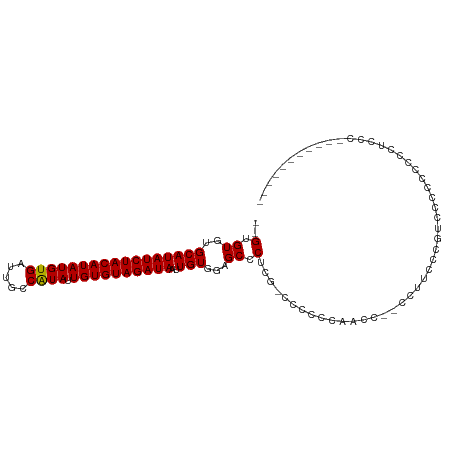
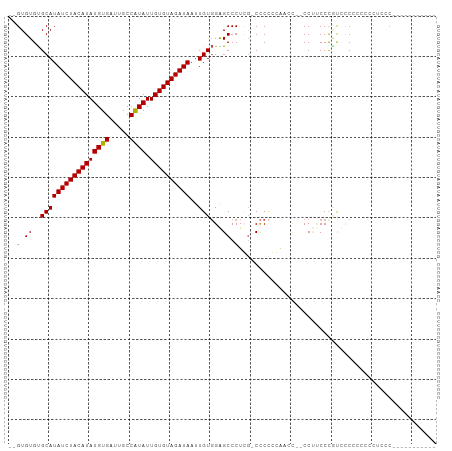
>dm3.chr3L 20567512 99 - 24543557 --GUGUGUGCAUAUCUACAUAUGUGAUUGCCAUAUUGUGUAGAUAAUUGUGGAGCCCUCG-CCCCUAAACC--CCUCCCCGCACCGCCUCUUCCCAACCCACUC-- --(((.((((.((((((((((((((.....)))).)))))))))).....((.((....)-).))......--.......))))))).................-- ( -25.40, z-score = -4.48, R) >droSim1.chr3L 19919848 99 - 22553184 --GUGUGUGCAUAUCUACAUAUGUGAUUGCCAUAUUGUGUAGAUAAUUGUGGAGCCCUCG-CCCCUCGAGC--CCCUCCCCUCCCACCCCUUCCCAUCACACUC-- --(((((.(((((((((((((((((.....)))).))))))))))..)))((((..((((-.....)))).--..))))..................)))))..-- ( -27.80, z-score = -4.86, R) >droYak2.chr3L 4212390 91 + 24197627 --GUGUGUGCAUAUCUACAUAUGUGAUUGCCAUAUUGUGUAGAUAAUUGUGGAGCCCUCGCCCCCGGAACC--CCUUCCCAUUCCCCUGCCUUUC----------- --......(((((((((((((((((.....)))).))))))))))...(.((.((....)).)))((((..--..))))........))).....----------- ( -19.90, z-score = -2.03, R) >droEre2.scaffold_4784 20261932 83 - 25762168 --GUGUGUGCAUAUCUACAUAUGUGAUUGCCGUAUUGUGUAGAUAAUUGUGGAGCCCUCG-CCCCGGAACC--CC-UCCCAUUCCUGUC----------------- --......(((((((((((((((((.....)))).))))))))))..)))((.((....)-).))((((..--..-.....))))....----------------- ( -15.60, z-score = -0.05, R) >droAna3.scaffold_13337 10920363 102 - 23293914 --GUGUGUGCAUAUCUACAUAUGUGAUUGCCAUAUUGUGUAGAUAAUUGUGGGGCCCUGGCCCCACCUCGU--CCUCCCUGCCACUGGCACCUCCGCCGCCAUCGC --..(((.(((((((((((((((((.....)))).))))))))))...(((((((....))))))).....--......)))))).(((......)))........ ( -39.50, z-score = -3.61, R) >droWil1.scaffold_180949 4130810 94 + 6375548 GUGUGUGCGCAUAUCUACAUAUGUGAUUGCCAUAUUGUGUAGAUAAUUGU-GGGCCCUGCCCUCUCUGCGCCGCCCGCCCGUCACCUGGGCACUC----------- (.(((.(((((((((((((((((((.....)))).))))))))))...(.-((((...)))).)..)))))))).)(((((.....)))))....----------- ( -38.60, z-score = -3.95, R) >consensus __GUGUGUGCAUAUCUACAUAUGUGAUUGCCAUAUUGUGUAGAUAAUUGUGGAGCCCUCG_CCCCCCAACC__CCUUCCCGUCCCCCCCCCUCCC___________ ....((..(((((((((((((((((.....)))).))))))))))..)))...))................................................... (-13.42 = -13.28 + -0.14)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:40:16 2011