
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 5,857,570 – 5,857,670 |
| Length | 100 |
| Max. P | 0.512999 |

| Location | 5,857,570 – 5,857,670 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 62.31 |
| Shannon entropy | 0.67040 |
| G+C content | 0.36874 |
| Mean single sequence MFE | -24.58 |
| Consensus MFE | -7.66 |
| Energy contribution | -7.80 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.52 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.512999 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 5857570 100 - 23011544 UUUGUAAACAGUUUGCCAU---GUUUUAUCGGCUGG---UUAGAUGUGAU--UUUCCGACUAUCUAGCUGAAAUUAUGAGCCCUUUGAAUUCG-CCUAAAAUUUAGAUU---------- ..............(((((---(.....((((((((---.(((.((.(..--...))).))).))))))))...)))).))..((((((((..-.....))))))))..---------- ( -18.70, z-score = -0.74, R) >droAna3.scaffold_12916 11268808 109 - 16180835 UGGACUAGCAUUUAAUCAUUUAAUUUAAUGGGCCAACAUUAAAAUGCAAUCAUUUUCAGAUUCGACGACUAAAGUGCAAGGAUAAUAGCUUCCCUUUCCGUUUCAGAUU---------- .(((...((((((..((......(((((((......))))))).((.((((.......))))))..))...))))))..(((........)))...)))..........---------- ( -17.00, z-score = 0.07, R) >droEre2.scaffold_4929 5941940 105 - 26641161 UCGGAAAACAGUUUGCCAU---GGUUUAUCAGCUGG---UUAGAGCAGAU--UUUCCAUCUGCCUAGCUGGAAUUAUGAGCACUUUCACUUUC-AGGAAUGUUCAAAUAACAAU----- ((.((((..((..((((((---(((((..(((((((---.....((((((--.....))))))))))))))))))))).)))..))...))))-..)).((((.....))))..----- ( -26.20, z-score = -1.28, R) >droYak2.chr2L 8983999 111 + 22324452 UCGAUAAACAGUUUGCCAU---GGUUUAUCAGCUGG---UUAGGAGAGAU--UUUCCGCCUGUCUAGCUGGAAUUAUGAGCCAUUUGAGCAGGUUUUUGGUUACAAAUAAAAUGGUAAU ............(((((((---((((((((((((((---.((((.(.(..--...)).)))).))))))).....)))))))((((((((.........))).)))))...))))))). ( -30.50, z-score = -1.76, R) >droSec1.super_5 3927420 102 - 5866729 UUGGUAAACAGUUUGCCAU---GGAUUAUAGGCUGG---UUAGAUGUGAU--UUUCCGACUAUCUAGCUGGAAUUAUGAGCCCAUUGAAUUUG-CCUAAAAUACUGACACU-------- ..((((((((((..(((((---((.((...((((((---.(((.((.(..--...))).))).)))))).)).))))).))..))))..))))-))...............-------- ( -23.20, z-score = -0.98, R) >droSim1.chr2L 5647792 102 - 22036055 UUGGUAAACAGUUUGCCAU---GGUUUAUCGGCUGG---AUAGAUGUCAU--UUUACGACUAUCUAGCUGGAAUUAUGAGCUCUUUGAAUUUG-CCUAAAUUACUAACACU-------- ..(((((((((...(((((---(((((..(((((((---((((.(((...--...))).))))))))))))))))))).))...)))..))))-))...............-------- ( -31.90, z-score = -4.48, R) >consensus UUGGUAAACAGUUUGCCAU___GGUUUAUCGGCUGG___UUAGAUGUGAU__UUUCCGACUAUCUAGCUGGAAUUAUGAGCCCUUUGAAUUUG_CCUAAAUUACAAAUA__________ ..((((((...))))))...........((((((((....(((.((.((.....)))).))).))))))))................................................ ( -7.66 = -7.80 + 0.14)
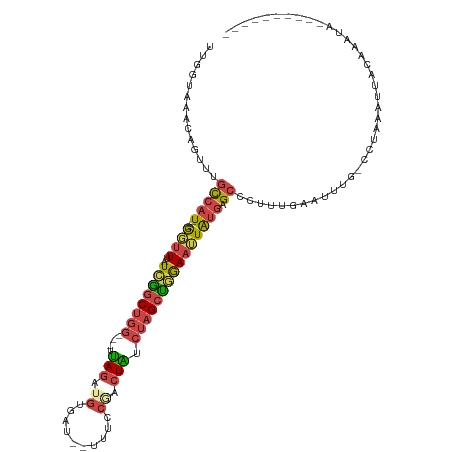


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:19:56 2011