
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 20,227,397 – 20,227,517 |
| Length | 120 |
| Max. P | 0.808679 |

| Location | 20,227,397 – 20,227,517 |
|---|---|
| Length | 120 |
| Sequences | 14 |
| Columns | 128 |
| Reading direction | reverse |
| Mean pairwise identity | 72.96 |
| Shannon entropy | 0.61251 |
| G+C content | 0.52529 |
| Mean single sequence MFE | -35.22 |
| Consensus MFE | -18.06 |
| Energy contribution | -17.84 |
| Covariance contribution | -0.21 |
| Combinations/Pair | 1.91 |
| Mean z-score | -1.04 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.808679 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 20227397 120 - 24543557 ------GUUCUCCGAAUGGGAACA--AAACAAUCUUACCCAGAAAAAUUGCGCCGGUUGCCACCAGCCAGGCAUCGCUGUGCGCUUCCUCCUCUUCGGGGAGCCGUACAAUCCGAACACCAUGCAGAC ------((((((((((.((((...--......(((.....)))......((((((((((((........))))..)))).))))....)))).))))))))))......................... ( -37.20, z-score = -1.79, R) >droSim1.chr3L 19597369 120 - 22553184 ------GUUCUCCGAGUGGGAACA--AAACAAUCUUACCCAGAAGAAUUGCGCCGGUUGCCACCAGCCAGGCAUCGCUGUGCGCUUCCUCCUCUUCGGGGAGCCGUACAACCCGAACACCAUGCAGAC ------((((((((((.((((...--......((((......))))...((((((((((((........))))..)))).))))....)))).))))))))))......................... ( -38.50, z-score = -1.47, R) >droSec1.super_11 131991 120 - 2888827 ------GUUCUCCGAGUGGGAACA--AAACAAUCUUACCCAGAAGAAUUGCGCCGGUUGCCACCAGCCAGGCAUCGCUGUGCGCUUCCUCCUCUUCGGGGAGCCGUACAAUCCGAACACCAUGCAGAC ------((((((((((.((((...--......((((......))))...((((((((((((........))))..)))).))))....)))).))))))))))......................... ( -38.50, z-score = -1.48, R) >droYak2.chr3L 3852577 120 + 24197627 ------GUUCUCCGAAUGGGAGCA--AAACAAUCUUACCCAGAAGAAUUGCGCCGGUUGCCACCAGCCAGGCAUCGCUGUGCGUUUCCUUCUCUUCGGAGAGCCGUAUAAUCCAAACACCAUGCAGAC ------((((((((((.(((((..--((((..((((......))))...((((((((.(((........)))))))..))))))))..))))))))))))))).((((............)))).... ( -37.90, z-score = -2.48, R) >droEre2.scaffold_4784 19951741 120 - 25762168 ------GUUCUCCGAAUGGGAACA--AAACAAUCUCACCCAGAAGAAUUGCGCCGGUUGCCACCAGCCAGGCAUCGCUGUGCGUUUCCUGCUCUUCGGGGAGCCGUAUAAUCCGAACACCAUGCAGAC ------((((((.....)))))).--.....................((((((((((((....))))).))).(((.(((((((((((((.....))))))).))))))...))).......)))).. ( -34.80, z-score = -0.45, R) >droAna3.scaffold_13337 6350294 120 - 23293914 ------AUUUUCCGAGUGGGAGCA--GAACAAUCUCACCCAAAAGAACUGUGCCGGUUGCCAUCAGCCGGGCAUUGCAGUGCGAUUUUUACUGUUUGGAGAACCGUAUAAUCCCAACACAAUGCAGAC ------...(((....((((.(.(--((....)))).))))...))).....(((((((....)))))))((((((...((.((((..(((.((((...)))).))).)))).))...)))))).... ( -32.20, z-score = -0.75, R) >dp4.chrXR_group6 2092076 120 - 13314419 ------CUUUUCCGAGUGGGAGCA--AAACAAUUUGACUCAGAAAAACUGCGCCGGCUGCCGACAGCCGGGCAUCGCAGUGCGCUUCCUUCUAUUCGGGGAGCCCUACAAUCCCAACACCAUGCAAAC ------.((..(((((((((((((--((....)))).))).(((..(((((((((((((....)))))))....))))))....)))...))))))))..)).......................... ( -40.30, z-score = -2.09, R) >droPer1.super_9 385303 120 - 3637205 ------CUUUUCCGAGUGGGAGCA--AAACAAUCUGACUCAGAAAAACUGCGCCGGCUGCCGACAGCCGGGCAUCGCAGUGCGCUUCCUUCUAUUCGGGGAGCCCUACAAUCCCAACACCAUGCAAAC ------.((..((((((((((((.--......((((...))))...(((((((((((((....)))))))....))))))..))))))....))))))..)).......................... ( -41.90, z-score = -2.52, R) >droWil1.scaffold_180698 3121367 120 - 11422946 ------AUUCUCUGAAUGGGAGCA--AAAUAAUCUGACACAAAAGAAUUGUGCCGGUUGCCAUCAGCCGGGUAUUGCUGUACGCUUUCUGCUAUUCGGAGAGCCCUACAAUCCCAAUACCAUGCAAAC ------..........((((((((--(..........(((((.....)))))(((((((....)))))))...))))((((.((((((((.....))))))))..)))).)))))............. ( -35.90, z-score = -2.41, R) >droVir3.scaffold_13049 18470272 120 - 25233164 ------UUUCUCUGAAUGGGUGCA--GAACAAUUUGACCCAAAAGAACUGCGCUGGCUGCCAUCAGCCGGGCAUUGCGGUGCGUUUCCUUCUGUUUGGGGAACCCUAUAAUCCAAACACCAUGCAGAC ------....(((...(((((.((--((....)))))))))..))).((((((((((((....)))))))((((....)))).........((((((((...........))))))))...))))).. ( -38.20, z-score = -1.19, R) >droMoj3.scaffold_6680 19271109 120 - 24764193 ------GUUCUCUGAGUGGGUGCA--GAACAACUUGACCCAGAAGAAUUGCGCUGGCUGCCAUCAACCAGGCAUUGCUGUUCGUUUCCUGCUAUUUGGGGAGCCCUACAAUCCAAACACCAUGCAGAC ------....((((....((((..--(((((.(((.......)))...(((.((((.((....)).)))))))....)))))(((((((.......))))))).............))))...)))). ( -30.70, z-score = 1.08, R) >droGri2.scaffold_15110 15386126 120 + 24565398 ------UUUCUCCGAGUGGGUGCA--GAACAAUUUAACCCAAAAGAACUGUGCUGGAUGCCAUCAGCCGGGCAUUGCCGUUCGUUUCCUUUUGUUUGGGGAACCCUAUAACCCAAGCACAAUGCAGAC ------.((((.....(((((...--..........)))))..))))((((..((((((((........))))))................((((((((...........))))))))))..)))).. ( -32.02, z-score = 0.07, R) >anoGam1.chr3L 38834162 92 - 41284009 ---------------------------------CUGCACCACCAGCACUGUGUCGCCUGCCAUCAGCCCGGCUUUGCCGCCCGGAUCGCGCUGCAGGGCAGCACCUACAAC---AGUGCCACGCUGGC ---------------------------------.....(((.(.(((((((((.(.(((((..(((((((((......).)))).....))))...)))))...).)).))---)))))...).))). ( -31.20, z-score = 1.28, R) >apiMel3.GroupUn 215955747 128 - 399230636 CUUAACGUUCUUAAAGAUAUACCAUUUACAGAAUAUAAAGGAAAGUCAUGUGCUGGAUGCCAACAGUUUAAAAUUCAUGCUAGAGUUCUAUUGUAUGGUCAACCAUACAAUGCUACUACAUUAGAAGG (((.(..((((((((.........)))).))))..).)))...((.((((.((((........))))........))))))..(((..((((((((((....))))))))))..)))........... ( -23.80, z-score = -0.33, R) >consensus ______GUUCUCCGAGUGGGAGCA__AAACAAUCUGACCCAGAAGAAUUGCGCCGGUUGCCAUCAGCCAGGCAUCGCUGUGCGCUUCCUUCUCUUCGGGGAGCCCUACAAUCCAAACACCAUGCAGAC ................((((.................))))......((((((((((((....))))).)))..........(((((((.......)))))))...................)))).. (-18.06 = -17.84 + -0.21)

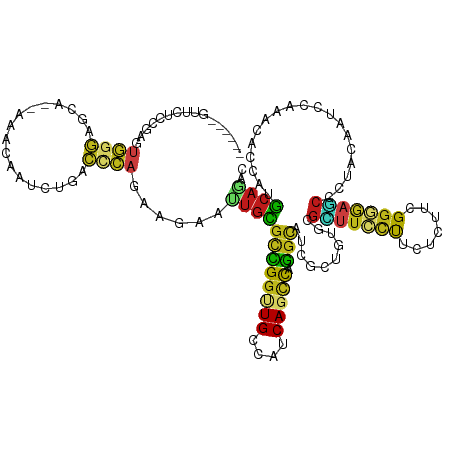

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:39:26 2011