
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 20,162,009 – 20,162,104 |
| Length | 95 |
| Max. P | 0.550134 |

| Location | 20,162,009 – 20,162,104 |
|---|---|
| Length | 95 |
| Sequences | 7 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 72.40 |
| Shannon entropy | 0.48368 |
| G+C content | 0.55857 |
| Mean single sequence MFE | -34.30 |
| Consensus MFE | -19.06 |
| Energy contribution | -20.41 |
| Covariance contribution | 1.35 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.15 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.550134 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
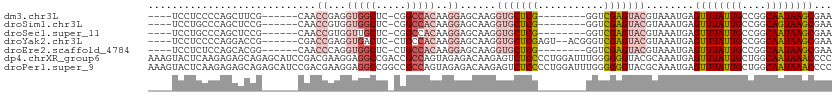
>dm3.chr3L 20162009 95 + 24543557 ----UCCUCCCCAGCUUCG------CAACCGAGGUGGCUC-CGGCCACAAGGAGCAAGGUGCUCG--------GGUCGAGUACGUAAAUGAGUUUAUUGCCGGCAAUAAGCGAA ----..((((...((((((------....))))))(((..-..)))....))))....(((((((--------...)))))))........((((((((....))))))))... ( -31.80, z-score = -0.62, R) >droSim1.chr3L 19449938 95 + 22553184 ----UCCUGCCCAGCUCCG------CAACCGUGGUGGCUC-CGGCCACAAGGAGCAAGGUGCUCG--------GGUCGAGUACGUAAAUGAGUUUAUUGCCGGCAGUAAGCGAA ----...(((...((((((------(....)).(((((..-..)))))..)))))...(((((((--------...)))))))))).....((((((((....))))))))... ( -35.30, z-score = -0.95, R) >droSec1.super_11 72374 95 + 2888827 ----UCCUGCCCAGCUCCG------CAACCGUGGUUGCUC-CGGCCACAAGGAGCAAGGUGCUCG--------GGUCGAGUACGUAAAUGAGUUUAUUGCCGGCAAUAAGCGAA ----...(((...((((((------((((....)))))..-.(....)..)))))...(((((((--------...)))))))))).....((((((((....))))))))... ( -33.70, z-score = -1.08, R) >droYak2.chr3L 3791448 101 - 24197627 ----UCCUCCCCAGGACCG------CGACCGAGGUGACUC-CUGCCACAAGGAGCAAGGUGCUCGAGU--ACGGGUCGAGUACGUAAAUGAGUUUAUUGCCGGCAAUAAGCGAA ----((((....))))..(------(((((....((.(((-((......))))))).)))((((((..--.....)))))).)))......((((((((....))))))))... ( -31.20, z-score = -0.34, R) >droEre2.scaffold_4784 19888759 95 + 25762168 ----UCCUCUCCAGCACGG------CAACCCAGGUGGCUC-CUGCCACAAGGAGCAAGGUGCUCG--------GGUCGAGUACGUAAAUGAGUUUAUUGCCGGCAAUAAGCGAA ----.........((.(((------(((.((..(((((..-..)))))..))......(((((((--------...))))))).............)))))))).......... ( -33.00, z-score = -1.19, R) >dp4.chrXR_group6 2035057 114 + 13314419 AAAGUACUCAAGAGAGCAGAGCAUCCGACGAAGGAGGCCGACCGCCAGUAGAGACAAGAGUCUCCCCUGGAUUUGGGGGGUACGCAAAUGAGUUUAUUGCUGGCAAUAAACCCC ...((((((.........(.((.(((......))).)))..((.((((..(((((....)))))..))))....)).))))))........((((((((....))))))))... ( -37.10, z-score = -2.06, R) >droPer1.super_9 328115 114 + 3637205 AAAGUACUCAAGAGAGCAGAGCAUCCGACGAAGGAGGCCGGCCGCCAGUAGAGACAAGAGUCUCCCCUGGAUUUGGGGGGUACGCAAAUGAGUUUAUUGCUGGCAAUAAACCCC ...((((((....(.((.(.((.(((......))).))).))).((((..(((((....)))))..)))).......))))))........((((((((....))))))))... ( -38.00, z-score = -1.79, R) >consensus ____UCCUCCCCAGCUCCG______CAACCGAGGUGGCUC_CGGCCACAAGGAGCAAGGUGCUCG________GGUCGAGUACGUAAAUGAGUUUAUUGCCGGCAAUAAGCGAA ............................((...(((((.....)))))..))......(((((((...........)))))))........((((((((....))))))))... (-19.06 = -20.41 + 1.35)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:39:19 2011