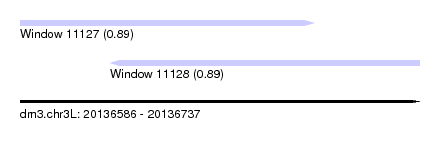
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 20,136,586 – 20,136,737 |
| Length | 151 |
| Max. P | 0.894673 |

| Location | 20,136,586 – 20,136,697 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 90.28 |
| Shannon entropy | 0.15211 |
| G+C content | 0.41626 |
| Mean single sequence MFE | -29.76 |
| Consensus MFE | -25.31 |
| Energy contribution | -25.38 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.09 |
| SVM RNA-class probability | 0.889084 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 20136586 111 + 24543557 CAGAAUUCAUAACUCAUUUCUUGAGUGUUUAAUGCACUCGGAAAAUG-CCGUCAAUGGAUUGGAGUUUUCCAACGAAGGAAACUCAUAAAGGUGCUCGGUGAGUCGUACUUU ........(..((((.(((((.((((((.....)))))))))))..(-(((.((........((((((.((......)))))))).......))..))))))))..)..... ( -29.36, z-score = -1.52, R) >droSim1.chr3L 19424433 104 + 22553184 CAGAAUUCAUAACUCAUUUCUUGAGUGUUUAAUGCACUCGGAAAAUA-CCGUCAAUGGAUUGGAGUUUCCCAACGAAGGAGACUCAUAAAGGUGCUCGGUGAGUC------- ...........((((.(((((.((((((.....)))))))))))..(-(((.((........((((((((.......)))))))).......))..)))))))).------- ( -31.26, z-score = -2.50, R) >droSec1.super_11 47271 104 + 2888827 CAGAAUUCAUAACUCAUUUCUUGAGUGUUUAAUGCACUCGGAAAAUA-CCGUCAUUGGAUUGGAGUUUCCCAACGAAGGAGACUCAUAAAGGUGCUCGGUGAGUC------- ...........((((.(((((.((((((.....)))))))))))..(-(((.((((......((((((((.......)))))))).....))))..)))))))).------- ( -33.50, z-score = -3.12, R) >droYak2.chr3L 3763176 104 - 24197627 CAGAAUUCAUAACUCAUGUCUUGAGUGUUUAAUGCACUUAGAAAAUAACCGUCAAUGGCUUGAAGUUUCCCAUCGACG-AGACUCAUAAAGGUGCACGGUGAGUC------- ...........((((...(((.((((((.....))))))))).....(((((((......(((.(((((........)-)))))))......)).))))))))).------- ( -24.90, z-score = -1.04, R) >consensus CAGAAUUCAUAACUCAUUUCUUGAGUGUUUAAUGCACUCGGAAAAUA_CCGUCAAUGGAUUGGAGUUUCCCAACGAAGGAGACUCAUAAAGGUGCUCGGUGAGUC_______ ...........((((((((((.((((((.....)))))))))))....(((...........((((((((.......))))))))...........))))))))........ (-25.31 = -25.38 + 0.06)


| Location | 20,136,620 – 20,136,737 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 76.56 |
| Shannon entropy | 0.33745 |
| G+C content | 0.46368 |
| Mean single sequence MFE | -26.43 |
| Consensus MFE | -19.84 |
| Energy contribution | -20.65 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.894673 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 20136620 117 - 24543557 GCAGCACUCAAAACUCCACCUUGGGAAAAGUACCUUGGGAAAAGUACGACUCACCGAGCACCUUUAUGAGUUUCCUUCGUUGGAAAACUCCAAUCCAUUGACGG-CAUUUUCCGAGUG ....(((((.....(((......)))...........((((((...(((((((..(((...)))..))))).......((((((....)))))).......)).-..))))))))))) ( -26.40, z-score = -0.22, R) >droSim1.chr3L 19424467 91 - 22553184 --------------------------ACUCCAACUUGGGAAAAGUUCGACUCACCGAGCACCUUUAUGAGUCUCCUUCGUUGGGAAACUCCAAUCCAUUGACGG-UAUUUUCCGAGUG --------------------------......((((((((((.(((((......)))))(((.....((((.((((.....)))).))))(((....)))..))-).)))))))))). ( -28.10, z-score = -2.46, R) >droSec1.super_11 47305 91 - 2888827 --------------------------ACUCCAACUUGGGAAAAGUACGACUCACCGAGCACCUUUAUGAGUCUCCUUCGUUGGGAAACUCCAAUCCAAUGACGG-UAUUUUCCGAGUG --------------------------......((((((((((.....((((((..(((...)))..)))))).((.((((((((.........)))))))).))-..)))))))))). ( -28.60, z-score = -2.79, R) >droYak2.chr3L 3763210 91 + 24197627 --------------------------AUUCCACCUUGAGUAAUGUACGACUCACCGUGCACCUUUAUGAGUCU-CGUCGAUGGGAAACUUCAAGCCAUUGACGGUUAUUUUCUAAGUG --------------------------....(((...(((((((....((((((.............)))))).-(((((((((...........)))))))))))))))).....))) ( -22.62, z-score = -1.51, R) >consensus __________________________ACUCCAACUUGGGAAAAGUACGACUCACCGAGCACCUUUAUGAGUCUCCUUCGUUGGGAAACUCCAAUCCAUUGACGG_UAUUUUCCGAGUG .................................(((((((((.....((((((..(((...)))..)))))).((.(((((((...........))))))).))...))))))))).. (-19.84 = -20.65 + 0.81)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:39:11 2011