
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 20,093,948 – 20,094,075 |
| Length | 127 |
| Max. P | 0.569856 |

| Location | 20,093,948 – 20,094,075 |
|---|---|
| Length | 127 |
| Sequences | 13 |
| Columns | 128 |
| Reading direction | reverse |
| Mean pairwise identity | 84.90 |
| Shannon entropy | 0.34258 |
| G+C content | 0.52606 |
| Mean single sequence MFE | -42.38 |
| Consensus MFE | -27.27 |
| Energy contribution | -27.29 |
| Covariance contribution | 0.02 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.569856 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
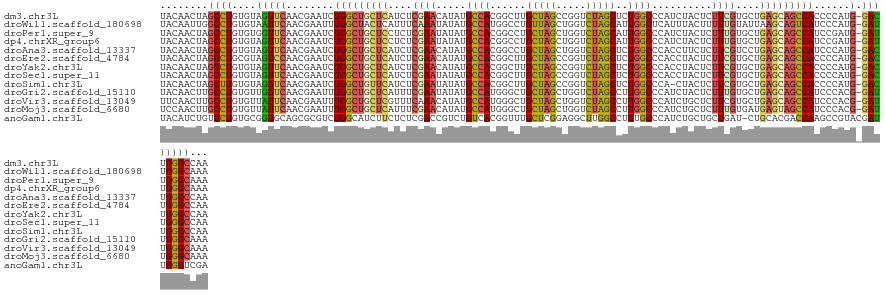
>dm3.chr3L 20093948 127 - 24543557 UACAACUAGCCUGUGUAGUUCAACGAAUCUGGCUGCUCAUCUCGAACAUAUGCCACGGCUUGCUAGCCGGUCUAGCUCUGGCCCAUCUACUCUUCGUGCUGAGCAGCCACCCCAUG-GACUGGGCCAA ........((((....((((((.......((((((((((.(.((((..........((((.(((((.....)))))...)))).........)))).).)))))))))).....))-))))))))... ( -43.41, z-score = -1.81, R) >droWil1.scaffold_180698 1257873 127 - 11422946 UACAAUUGGCCUGUGUAAUUCAACGAAUUUGGCUACUCAUUUCAAAUAUAUGCCAUGGCCUGUUAGCUGGUCUAGCAUUGGCUCAUUUACUUUUUGUAUUAAGCAGUCAUCCCAUG-GAUUGGGCAAA ......(((((.....(((((...))))).)))))...............((((((((((((((((.....))))))..))).))).................(((((........-))))))))).. ( -26.10, z-score = 1.19, R) >droPer1.super_9 2977898 127 - 3637205 UACAACUAGCCUGUGUGGUUCAACGAAUCUGGCUGCUCCUCUCGAAUAUAUGCCACGGCCUGCUAGCUGGUCUAGCAUUGGCCCAUCUACUCUUUGUGCUGAGCAGCCAUCCGAUG-GAUUGGGCAAA ........((((....((((((.((....(((((((((..(.((((..........((((((((((.....))))))..)))).........)))).)..)))))))))..)).))-))))))))... ( -42.31, z-score = -1.33, R) >dp4.chrXR_group6 4677239 127 - 13314419 UACAACUAGCCUGUGUAGUUCAACGAAUCUGGCUGCUCCUCUCGAAUAUAUGCCACGGCCUGCUAGCUGGUCUAGCAUUGGCCCAUCUACUCUUUGUGCUGAGCAGCCAUCCGAUG-GAUUGGGCAAA ...........(((.(((((((.((....(((((((((..(.((((..........((((((((((.....))))))..)))).........)))).)..)))))))))..)).))-))))).))).. ( -42.71, z-score = -1.65, R) >droAna3.scaffold_13337 18253956 127 + 23293914 UACAACUAGCCUGUGUAGUUCAACGAAUCUGGCUGCUCAUCUCGAACAUAUGCCACGGCCUGCUAGCUGGUCUAGCUCUGGCCCACCUUCUCUUCGUCCUGAGCAGCCAUCCCAUG-GACUGGGCCAA ........((((....((((((..((...((((((((((...((((..........((((.(((((.....)))))...)))).........))))...))))))))))))...))-))))))))... ( -44.41, z-score = -2.50, R) >droEre2.scaffold_4784 19831754 127 - 25762168 UACAACUAGCCUGCGUAGUCCAACGAAUCUGGCUGCUCAUCUCGAACAUAUGCCACGGCUUGCUAGCCGGUCUAGCUCUGGCCCACCUACUCUUCGUGCUGAGCAGCCACCCCAUG-GACUGGGCCAA ............((.(((((((.......((((((((((.(.((((..........((((.(((((.....)))))...)))).........)))).).)))))))))).....))-))))).))... ( -47.31, z-score = -2.86, R) >droYak2.chr3L 3728798 127 + 24197627 UACAACUAGCCUGUGUAGUUCAACGAAUCUGGCUGCUCAUCUCGAACAUAUGCCACGGCUUGCUAGCCGGUCUAGCUCUGGCCCACCUACUCUUCGUGCUGAGCAGCCACCCCAUG-GACUGGGCCAA ........((((....((((((.......((((((((((.(.((((..........((((.(((((.....)))))...)))).........)))).).)))))))))).....))-))))))))... ( -43.41, z-score = -1.76, R) >droSec1.super_11 9960 127 - 2888827 UACAACUAGCCUGUGUAGUUCAACGAAUCUGGCUGCUCAUCUCGAAUAUAUGCCACGGCUUGCUAGCCGGUCUAGCUCUGGCCCACCUACUCUUCGUGCUGAGCAGCCACCCCAUG-GACUGGGCCAA ........((((....((((((.......((((((((((.(.((((..........((((.(((((.....)))))...)))).........)))).).)))))))))).....))-))))))))... ( -43.41, z-score = -1.81, R) >droSim1.chr3L 19388745 126 - 22553184 UACAACUAGCUUGUGUAGUUCAACGAAUCUGGCUGUUCAUCUCGAAUAUAUGCCACGGCUUGCUAGCCGGUCUAGCUCUGGCCCA-CUACUCUUCGUGCUGAGCAGCCACCCCAUG-GACUGGGCCAA ............((.(((((((.......((((((((((.(.((((..........((((....))))((((.......))))..-......)))).).)))))))))).....))-))))).))... ( -40.40, z-score = -1.22, R) >droGri2.scaffold_15110 15202259 127 + 24565398 UACAACUUGCCUGUGUUGUUCAACGAAUUUGGCUGCUCAUUUCGAAUAUAUGCCAUGGGCUGCUAGCUGGUCUAGCCUUGGCCCAUCUACUCUUUGUGCUGAGCAGCCAUCCCACG-GAUUGGGCAAA ......((((((.((((....))))....((((((((((...((((........((((((((((((.....)))))...)))))))......))))...)))))))))).......-....)))))). ( -43.64, z-score = -2.04, R) >droVir3.scaffold_13049 18280459 127 - 25233164 UUCAACUUGCCUGUGUUAUUCAACGAAUUUGGCUGCUCGUUUCAAACAUAUGCCAUGGGCUGCUAGCUGGUCUAGCCUUGGCCCAUCUGCUCUUCGUGCUGAGCAGCCAUCCCACG-GAUUGGGCAAA ......((((((.................((((((((((...((.......((.((((((((((((.....)))))...)))))))..))......)).))))))))))(((...)-))..)))))). ( -43.32, z-score = -1.70, R) >droMoj3.scaffold_6680 19062510 127 - 24764193 UCCAACUUGCCUGUGUUAUUCAACGAAUUUGGCUGCUCAUUUCGAACAUAUGCCAUGGGCUGCUAGCUGGUCUAGCCUUGGCCCAUCUGCUCUUUGUGAUGAGUAGCCAUCCCACG-GAUUGGGCAAA ......((((((.................((((((((((((.((((.....((.((((((((((((.....)))))...)))))))..))..)))).))))))))))))(((...)-))..)))))). ( -46.70, z-score = -3.00, R) >anoGam1.chr3L 6252107 127 + 41284009 UACAUCUGUCCUGUGCGGUGCAGCGCGUCUGGCAUCUUCUCUCGACCGUCUGUCACGGUUUGCUCGGAGGCUUGGCCUUUGCCCAUCUGCUGCUGAU-CUGCACGACCAAGCCGUACGAUUGGGUCGA ............((((((.(((((((.(((((((.........((((((.....))))))))).)))).))..(((....))).....)))))....-))))))((((....(....)....)))).. ( -43.80, z-score = -0.18, R) >consensus UACAACUAGCCUGUGUAGUUCAACGAAUCUGGCUGCUCAUCUCGAACAUAUGCCACGGCCUGCUAGCUGGUCUAGCUUUGGCCCAUCUACUCUUCGUGCUGAGCAGCCAUCCCAUG_GAUUGGGCCAA ........((((.(((......)))....(((((((((....((((.....((((......(((((.....)))))..))))..........))))....)))))))))............))))... (-27.27 = -27.29 + 0.02)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:39:03 2011