| Sequence ID | dm3.chr3L |
|---|---|
| Location | 19,570,111 – 19,570,167 |
| Length | 56 |
| Max. P | 0.953579 |

| Location | 19,570,111 – 19,570,167 |
|---|---|
| Length | 56 |
| Sequences | 6 |
| Columns | 56 |
| Reading direction | forward |
| Mean pairwise identity | 64.01 |
| Shannon entropy | 0.67660 |
| G+C content | 0.41701 |
| Mean single sequence MFE | -14.18 |
| Consensus MFE | -5.35 |
| Energy contribution | -5.88 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.953579 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
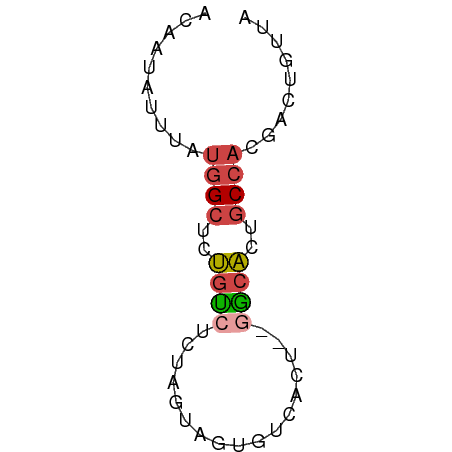
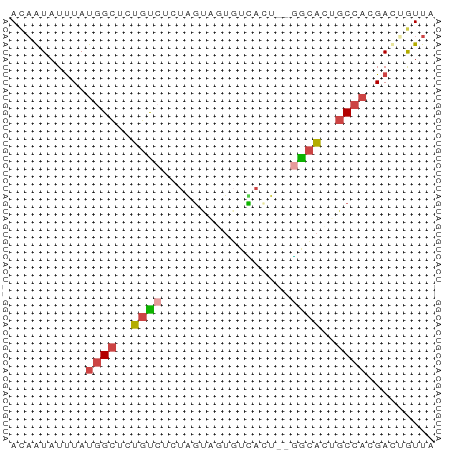
>dm3.chr3L 19570111 56 + 24543557 ACAAUAUUUAUGGCUCUGUGUUUAGUUGUUUUACUCUAGCACUGCCACGAUUGUUA ..((((.(..((((...(((((.(((......)))..))))).))))..).)))). ( -14.70, z-score = -2.97, R) >droYak2.chr3L 3223868 54 - 24197627 ACAAUAUUUAUGGCUGUGUCUUUAGUUGUGUUACU--GGCACGGCCACGAUUGUUA ..((((.(..((((((((((...(((......)))--))))))))))..).)))). ( -19.50, z-score = -3.73, R) >droAna3.scaffold_13337 5974556 53 - 23293914 AUAAUAUUUAUGGCUCUGUCUCUAGUUGUGUCACU--GGCACGGCCACGAUUGUU- ..((((.(..(((((.((((...(((......)))--)))).)))))..).))))- ( -12.70, z-score = -1.32, R) >dp4.chrXR_group8 7645745 54 + 9212921 AUAAUAUUUAUGGCUCUGCCACUGGCACUGGCAAU--GGCACUGCCAAGACGGUUC ..........((((..(((((...((....))..)--))))..))))......... ( -16.70, z-score = -1.79, R) >droPer1.super_48 424037 54 + 581423 AUAAUAUUUAUGGCUCUGCCACUGGCACUGGCAAU--GGCACUGCCAAGACGGUUC ..........((((..(((((...((....))..)--))))..))))......... ( -16.70, z-score = -1.79, R) >triCas2.ChLG10 8364025 56 + 8806720 ACUGUAUCUACCGUGGGAUUUUUUUUAAUUUAUGCUCUAGUUAUCCAUGACAAUUA ...........((((((........(((((........)))))))))))....... ( -4.80, z-score = 0.50, R) >consensus ACAAUAUUUAUGGCUCUGUCUCUAGUAGUGUCACU__GGCACUGCCACGACUGUUA ..........((((..((((...((........))..))))..))))......... ( -5.35 = -5.88 + 0.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:38:08 2011