
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 19,565,149 – 19,565,245 |
| Length | 96 |
| Max. P | 0.509555 |

| Location | 19,565,149 – 19,565,245 |
|---|---|
| Length | 96 |
| Sequences | 11 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 59.61 |
| Shannon entropy | 0.82852 |
| G+C content | 0.34492 |
| Mean single sequence MFE | -10.35 |
| Consensus MFE | -5.83 |
| Energy contribution | -5.83 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | 0.30 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.509555 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
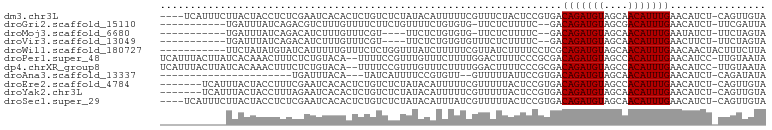
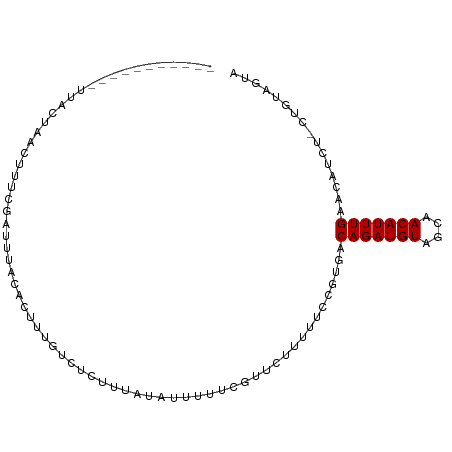
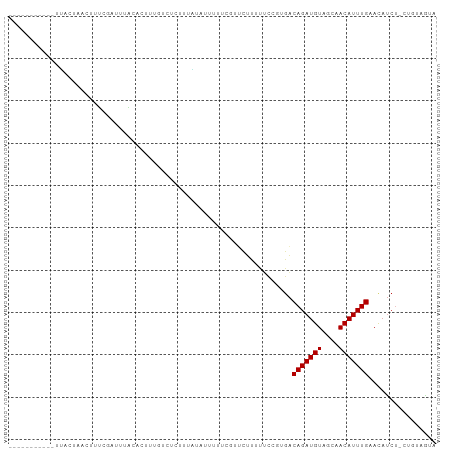
>dm3.chr3L 19565149 96 - 24543557 ----UCAUUUCUUACUACCUCUCGAAUCACACUCUGUCUCUAUACAUUUUUCGUUUCUACUCCGUGACAGAUGUAGCAACAUUUGAACAUCU-CAGUUGUA ----........................(((.((((((...........................))))))))).(((((...(((.....)-))))))). ( -11.63, z-score = -0.13, R) >droGri2.scaffold_15110 23889274 86 - 24565398 -----------UGAUUUAUCAGACGUCUUUGUUUUCUUCUGUUUUCUGUGUG-UUCUCUUUUC--GACAGAUGUAGCGACAUUUGAACAUCU-UUCGAUUA -----------.(((...(((((.(((...(((....((((((....(.(..-..).).....--))))))...)))))).)))))..))).-........ ( -11.50, z-score = 1.15, R) >droMoj3.scaffold_6680 10546329 82 + 24764193 -----------UGAUUUAUCAGACAUCUUUGUUUCGU----UUCUCUGUGUG-UUCUCUUUUC--GACAGAUGUAGCAACAUUUGAAUAUCU-UUCUAGUA -----------.(((...(((((((....))))..((----(.((..((.((-((........--)))).))..)).)))...)))..))).-........ ( -9.20, z-score = 1.29, R) >droVir3.scaffold_13049 13144103 83 - 25233164 -----------UGAUUUAUCAGACAUCUUUGUUUCGU----UUCUCUGUGUGUUUCUCUUUUC--GACAGAUGUAGCAACAUUUGAACUUCU-UUCUAGUA -----------.((..(((((((..............----...)))).)))..)).......--..(((((((....))))))).......-........ ( -8.73, z-score = 1.51, R) >droWil1.scaffold_180727 850703 90 - 2741493 -----------UUCUAUAUGUAUCAUUUUUGUUUCUCUGGUUUAUCUUUUUCGUUAUCUUUUCCUCGCAGAUGUAGCAACAUUUGAACAACUACUUUCUUA -----------.......(((.(((...(((((.(((((............................)))).).)))))....))))))............ ( -6.79, z-score = 0.21, R) >droPer1.super_48 420107 98 - 581423 UCAUUUACUUAUCACAAACUUUCUCUGUACA--UUUUCCGUUUGUUUCUUUUGGACUUUUCCCGCGACAGAUGUAGCCACAUUUGAACAUCC-UUGUAAUA ....((((...((((((((............--......)))))).......(((....)))...))(((((((....))))))).......-..)))).. ( -10.67, z-score = -0.02, R) >dp4.chrXR_group8 7641792 98 - 9212921 UCAUUUACUUAUCACAAACUUUCUCUGUACA--UUUUCCGUUUGUUUCUUUUGGACUUUUCCCGCGACAGAUGUAGCCACAUUUGAACAUCC-UUGUAAUA ....((((...((((((((............--......)))))).......(((....)))...))(((((((....))))))).......-..)))).. ( -10.67, z-score = -0.02, R) >droAna3.scaffold_13337 5968875 73 + 23293914 ----------------------UGAUUUACA---UAUCAUUUUCCGUGUU--GUUUUUAUUCCGUGACAGAUGUAGCAACAUUUGAACAUCU-CAGAUAUA ----------------------........(---((((.......(((((--(((..((((........)))).)))))))).(((.....)-))))))). ( -10.60, z-score = -0.28, R) >droEre2.scaffold_4784 19362508 93 - 25762168 -------UCAUUUACUACCUUUCGAAUCACACUCUGUCUCUAUACAUUUUUCGUUUUUACUCCGUGACAGAUGUAGCCACAUUUGAACAUCU-CAGUUGUA -------.....................(((.((((((...........................)))))))))....(((.((((.....)-))).))). ( -10.83, z-score = -0.14, R) >droYak2.chr3L 3216961 93 + 24197627 -------UCAUUUACUACCUUUAGAAUCACACUCUGUCUCUAUACAUUUUUCGUUUUUACUCCGUGACAGAUGUAGCAACAUUUGAACAUCU-CAGUUGUA -------.....................(((.((((((...........................))))))))).(((((...(((.....)-))))))). ( -11.63, z-score = -0.16, R) >droSec1.super_29 269195 96 - 700605 ----UCAUUUCUUACUACCUCUCGAAUCACACUCUGUCUCUAUACAUUUAUCGUUUUUACUCCGUGACAGAUGUAGCAACAUUUGAACAUCU-CAGUUGUA ----........................(((.((((((...........................))))))))).(((((...(((.....)-))))))). ( -11.63, z-score = -0.08, R) >consensus ___________UUACUAACUUUCGAUUUACACUUUGUCUCUUUAUAUUUUUCGUUCUUUUUCCGUGACAGAUGUAGCAACAUUUGAACAUCU_CUGUAGUA ...................................................................(((((((....)))))))................ ( -5.83 = -5.83 + -0.00)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:38:07 2011