
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 19,450,169 – 19,450,282 |
| Length | 113 |
| Max. P | 0.615215 |

| Location | 19,450,169 – 19,450,282 |
|---|---|
| Length | 113 |
| Sequences | 8 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 65.75 |
| Shannon entropy | 0.65969 |
| G+C content | 0.52996 |
| Mean single sequence MFE | -23.81 |
| Consensus MFE | -11.60 |
| Energy contribution | -11.53 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.19 |
| Mean z-score | -0.68 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.615215 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 19450169 113 - 24543557 CUCUCCCCCUCCCACCUUCUCGGCCACCCUAAACACGCCCCUUUUCCACCUGACACGGAUGCACA----AUGCGCACGUGUUCAAUAACAAUUGUGUUGCAUACUUUAGUGCGCCGG ....................((((.......((((((..............((((((..(((...----..)))..))))))..........))))))((((......)))))))). ( -24.45, z-score = -1.95, R) >droSim1.chr3L 18780563 108 - 22553184 -----CCCCCGCCCCCUUCUCUGCCACCCUAAACACGCCCCUUUUCCACCUGACACGGAUGCACA----AUGCGCACGUGUUCAAUAACAAUUGUGUUGCAUACUUUAGUGCGCCGG -----..((.((...................((((((..............((((((..(((...----..)))..))))))..........))))))((((......)))))).)) ( -20.55, z-score = -0.70, R) >droSec1.super_29 151716 108 - 700605 -----CGCCCCACCCCCUCUCUUCCACCCUAAGCACGCCCCUUUUCCACCUGACACGGAUGCACA----AUGCGCACGUGUUCAAUAACAAUUGUGUUGCAUACUUUAGUGCGCCGG -----......................((...((((...............((((((..(((...----..)))..))))))...(((((....))))).........))))...)) ( -20.00, z-score = -0.44, R) >droYak2.chr3L 3105971 102 + 24197627 -----------CCCCAGCCCCACCCCCUGCAUCCACGACCCUUUUCCACCUGACACGGAUGCACA----AUGCGCACGUGUUCAAUAACAAUUGUGUUGCAUACUUUAGUGCGCCGG -----------........((......(((((((......................)))))))..----..((((((((((.(((((.(....)))))).))))....)))))).)) ( -23.85, z-score = -1.62, R) >droAna3.scaffold_13337 5263913 102 + 23293914 -----------CCCUAUUCCGAUCCCACUUUGCUUUUUUUUUGUUCCUUCUGACACGGAUGCACA----ACGCGCACGUGUUCAAUAACAAUUGUGUUGCAUACUUUAGUGCGCCAG -----------...................(((....((..((((......))))..)).)))..----..((((((((((.(((((.(....)))))).))))....))))))... ( -18.40, z-score = -0.54, R) >dp4.chrXR_group8 7531268 103 - 9212921 --------------UCCUCUCCUGCUCUUGCUCCUGCUCCUUAUCCGGCUCCACAUGGCUGCAGAGGGCGCGUGCACGUGUUCAAUAACAAUUGUGUUGCAUACUUUAGUGCAGCAG --------------.......((((....((....)).........((((......)))))))).(((((((....)))))))...........((((((((......)))))))). ( -28.10, z-score = 0.04, R) >droPer1.super_48 309189 97 - 581423 -------------------UCCU-CUCCUGCUCCUGCUCCUUAUCCGGCUCCACAUGGCUGCAGAGGGCGCGUGCACGUGUUCAAUAACAAUUGUGUUGCAUACUUUAGUGCAGCAG -------------------....-...((((....((.((((...(((((......)))))...)))).)).(((((((((.(((((.(....)))))).))))....))))))))) ( -28.00, z-score = -0.42, R) >droGri2.scaffold_15110 23724184 92 - 24565398 ----------------------GCCGCUGCGGCUCGAUGCAAGUUGCAACCUGCUUGCAUUUGCUG---GCUGCUACGUGUUCAAUAACCAUUGUGUUGCAUACUUUAGUGUGAAGG ----------------------..((((((((((.(((((((((........)))))))))....)---)))))...((((.(((((.......))))).))))...))))...... ( -27.10, z-score = 0.15, R) >consensus ___________CCCCCUUCUCAUCCACCCUAACCACGUCCCUUUUCCACCUGACACGGAUGCACA____ACGCGCACGUGUUCAAUAACAAUUGUGUUGCAUACUUUAGUGCGCCGG .......................................................................((((((((((.(((((.(....)))))).))))....))))))... (-11.60 = -11.53 + -0.08)

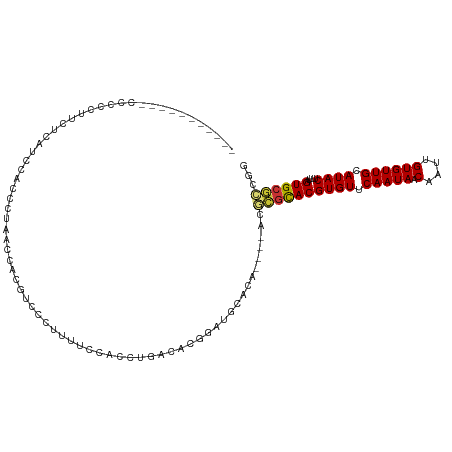

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:37:49 2011