| Sequence ID | dm3.chr3L |
|---|---|
| Location | 19,344,628 – 19,344,721 |
| Length | 93 |
| Max. P | 0.978198 |

| Location | 19,344,628 – 19,344,721 |
|---|---|
| Length | 93 |
| Sequences | 10 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 73.91 |
| Shannon entropy | 0.52097 |
| G+C content | 0.42521 |
| Mean single sequence MFE | -25.52 |
| Consensus MFE | -14.27 |
| Energy contribution | -15.16 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.99 |
| SVM RNA-class probability | 0.978198 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 19344628 93 + 24543557 --------CAUUCCGCUCCAUUCCG---CCUAGGCCCUGAUUAUUGUUGACUGUGUUAAUAAUGA-AAAAGACAGUUGCAAACAGCUGUGGCUUGGCUGAAGAAA------ --------............(((.(---((.(((((((..(((((((((((...)))))))))))-...))(((((((....))))))))))))))).)))....------ ( -29.10, z-score = -3.19, R) >droSim1.chr3L 18669891 93 + 22553184 --------CAUUCCGCUCCAUUCCG---CCUAGGCCCUGAUUAUUGUUGACUGUGUUAAUAAUGA-AAAAGACAGUUGCAAACAGCUGUGGCUUGGCUGAAGAAA------ --------............(((.(---((.(((((((..(((((((((((...)))))))))))-...))(((((((....))))))))))))))).)))....------ ( -29.10, z-score = -3.19, R) >droSec1.super_29 46145 93 + 700605 --------CAUUCCGCUCCAUUCCG---CCUAGGCCCUGAUUAUUGUUGACUGUGUUAAUAAUGA-AAAAGACAGUUGCAAACAGCUGUGGCUUGGCUGAAGAAA------ --------............(((.(---((.(((((((..(((((((((((...)))))))))))-...))(((((((....))))))))))))))).)))....------ ( -29.10, z-score = -3.19, R) >droYak2.chr3L 2979122 101 - 24197627 -----CUACAUUCCGCACCAUUCCG---CCUAGGCCCUGAUUAUUGUUGACUGUGUUAAUAAUGA-AAAAGACAGUUGCAAACAGCUGUGGCUUGGCUGAAGAAACUGAA- -----...............(((.(---((.(((((((..(((((((((((...)))))))))))-...))(((((((....))))))))))))))).))).........- ( -29.10, z-score = -2.82, R) >droAna3.scaffold_13337 5177606 95 - 23293914 --------CAUUCCGUUCCAUUCCAGUUCCUAGGCCCUGAUUAUUGUUGACUGUGUUAAUAAUGG-AAAAGACAGUUGCAAACAGCUGUGGCUUGACUGAAGUA------- --------...............(((((...(((((((..(((((((((((...)))))))))))-...))(((((((....))))))))))))))))).....------- ( -24.70, z-score = -1.57, R) >dp4.chrXR_group8 7430501 101 + 9212921 CAUUCCGACAUUCCAUUCCAUUCCG---CCUAGGCCCUGAUUAUUGUUGACUGUGUUAAUAAUGG-AAAAGACAGUUGCAAACAGCUGUGGCUUGGCUUGACUGG------ .................(((.((.(---((.(((((((..(((((((((((...)))))))))))-...))(((((((....)))))))))))))))..)).)))------ ( -28.80, z-score = -1.82, R) >droPer1.super_48 220342 101 + 581423 CAUUCCGACAUUCCAUUCCAUUCCG---CCUAGGCCCUGAUUAUUGUUGACUGUGUUAAUAAUGG-AAAAGACAGUUGCAAACAGCUGUGGCUUGGCUUGACUGG------ .................(((.((.(---((.(((((((..(((((((((((...)))))))))))-...))(((((((....)))))))))))))))..)).)))------ ( -28.80, z-score = -1.82, R) >droWil1.scaffold_180727 634366 94 + 2741493 -------GUCUCUUUUUCGCUUUCCCUAGGCUCUCUCUGAUUAUUGUUGGCUGUGUUAAUAAUGGAAAAAGACAGCUGCAAACAGAUGUGACCCAAAGGCA---------- -------(((((((((((((((.....))))..........((((((((((...)))))))))))))))))(((.(((....))).))).......)))).---------- ( -20.90, z-score = -0.14, R) >droVir3.scaffold_13049 12932562 74 + 25233164 ----------------------------UCUAGGCAGUGAUUAUUGUUGACUGUGUUAAUAAUG--AAAAGACAGUUACAAACAGCUGUGGCUUCUGUGAAGCA------- ----------------------------.....(((((.....((((.((((((.((.......--..)).))))))))))...))))).(((((...))))).------- ( -18.70, z-score = -1.13, R) >droGri2.scaffold_15110 23603621 81 + 24565398 ----------------------------UCUAGUCAGUGAUUAUUGUUGACUGUGUUAAUAAUG--AAAAGACAUUUCUAAACAGCUGUGGCUUCUGUAAGGCAGGAGAAA ----------------------------......((((..(((((((((((...))))))))))--)..(((....))).....))))...((((((.....))))))... ( -16.90, z-score = -0.48, R) >consensus ________CAUUCCGCUCCAUUCCG___CCUAGGCCCUGAUUAUUGUUGACUGUGUUAAUAAUGA_AAAAGACAGUUGCAAACAGCUGUGGCUUGGCUGAAGAAA______ ...............................(((((....(((((((((((...)))))))))))......(((((((....))))))))))))................. (-14.27 = -15.16 + 0.89)
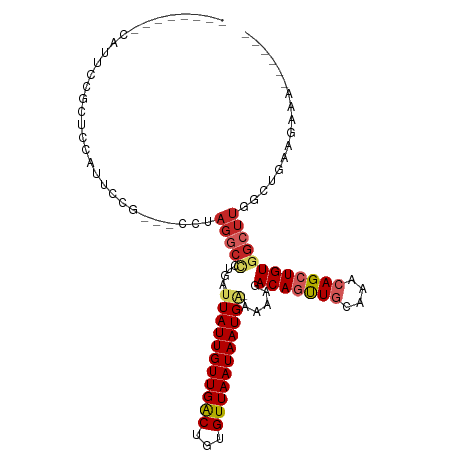


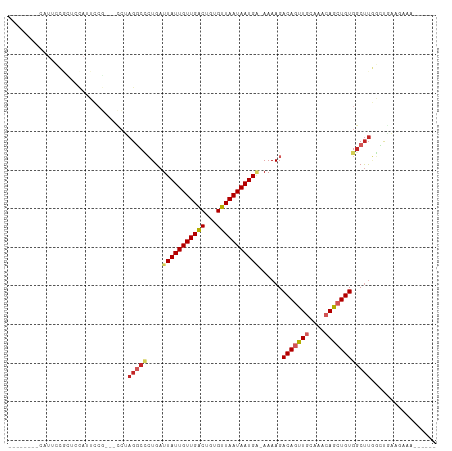
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:37:33 2011