
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 19,282,339 – 19,282,445 |
| Length | 106 |
| Max. P | 0.830437 |

| Location | 19,282,339 – 19,282,445 |
|---|---|
| Length | 106 |
| Sequences | 7 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 75.82 |
| Shannon entropy | 0.43491 |
| G+C content | 0.40422 |
| Mean single sequence MFE | -30.03 |
| Consensus MFE | -14.99 |
| Energy contribution | -15.29 |
| Covariance contribution | 0.29 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.830437 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
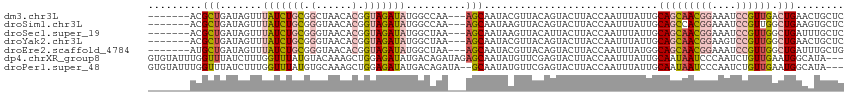
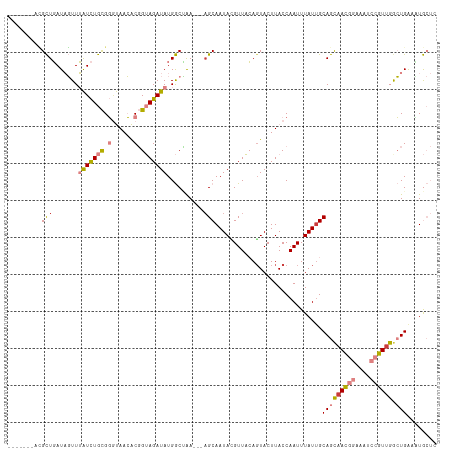
>dm3.chr3L 19282339 106 - 24543557 -------ACGCUGAUAGUUUAUCUGCGGCUAACACGGUAGAUAUGGCCAA---AGCAAUACGUUACAGUACUUACCAAUUUAUUGCAGCAACGGAAAUCCGUUGACUGAACUGCUC -------.(((.((((...)))).)))((.....((((((((.(((...(---((...(((......)))))).)))))))))))(((((((((....)))))).)))....)).. ( -26.70, z-score = -1.34, R) >droSim1.chr3L 18591938 106 - 22553184 -------ACGCUGAUAGUUUAUCUGCGGGUAACACGGUAGAUAUGGCCAA---AGCAAUAAGUUACAGUACUUACCAAUUUAUUGCAGCCACGGAAAUCCGUUGGCUGAAGUGCUC -------..(((.......(((((((.(......).))))))).......---)))..........(((((((..(((....)))(((((((((....))).))))))))))))). ( -33.84, z-score = -2.64, R) >droSec1.super_19 1236170 106 - 1257711 -------ACGCUGAUAGUUUAUCUGCGGGUAACACGGUAGAUAUGGCUAA---AGCAAUAAGUUACAUUACUUACCAAUUUAUUGCAGCAACGGAAAUCCGUUGGCUGAUUUGCUC -------..(((..((((((((((((.(......).))))))).))))).---.((((((((((............)))))))))))))(((((....)))))(((......))). ( -31.80, z-score = -3.00, R) >droYak2.chr3L 2920655 106 + 24197627 -------ACGCUGAUAGUUUAUCUGCGGGUAACACGGUAGAUAUGGCUAA---AGCAAUACGUUACAGUACUUACCAAUUUAUUGCAGCAACGGAAGUCCGUUGGCUGAACUGCUC -------..(((..((((((((((((.(......).))))))).))))).---))).........((((......(((....)))(((((((((....))))).)))).))))... ( -31.10, z-score = -1.78, R) >droEre2.scaffold_4784 19094265 106 - 25762168 -------AUGCUGAUAGUUUAUCUGCGGGUAACACGGUAGAUAUGGCUAA---AGCAAUACGUUACAGUACUUACCAAUUUAUGGCAGCAACGGAAAUCCGUUGGCUGAUUUGCUG -------.((((..((((((((((((.(......).))))))).))))).---))))........(((((....(((.....)))(((((((((....))))).))))...))))) ( -35.00, z-score = -3.23, R) >dp4.chrXR_group8 7370178 113 - 9212921 GUGUAUUUGGUUUAUCUUUGGUUUAUGUACAAAGCUGGAGAUAUGACAGAUAGAGCAAUAUGUUCGAGUACUUACCAAUUUAUUGCAAUAAUCCCAAUCUGUUGAAUGGCAUA--- .(.((((((...((((((..((((.......))))..))))))...)))))).)((((((.(((.(.((....)))))).)))))).......(((.((....)).)))....--- ( -26.10, z-score = -1.11, R) >droPer1.super_48 160273 111 - 581423 GUGUAUUUGGUUUAUCUUUGGUUUAUGUGCAAAGCUGGAGAUAUGACAGAUA--GCAAUAUGUUCGAGUACUUACCAAUUUAUUGCAAUAAUCCCAAUCUGUUGAAUGGCAUA--- ...((((((...((((((..((((.......))))..))))))...))))))--((((((.(((.(.((....)))))).)))))).......(((.((....)).)))....--- ( -25.70, z-score = -0.88, R) >consensus _______ACGCUGAUAGUUUAUCUGCGGGUAACACGGUAGAUAUGGCUAA___AGCAAUACGUUACAGUACUUACCAAUUUAUUGCAGCAACGGAAAUCCGUUGGCUGAAAUGCUC .........(((.......(((((((.(......).))))))).))).......((((((.(((............))).))))))..((((((....))))))............ (-14.99 = -15.29 + 0.29)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:37:25 2011