
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 18,845,504 – 18,845,622 |
| Length | 118 |
| Max. P | 0.772944 |

| Location | 18,845,504 – 18,845,622 |
|---|---|
| Length | 118 |
| Sequences | 9 |
| Columns | 129 |
| Reading direction | forward |
| Mean pairwise identity | 68.72 |
| Shannon entropy | 0.65001 |
| G+C content | 0.51128 |
| Mean single sequence MFE | -23.35 |
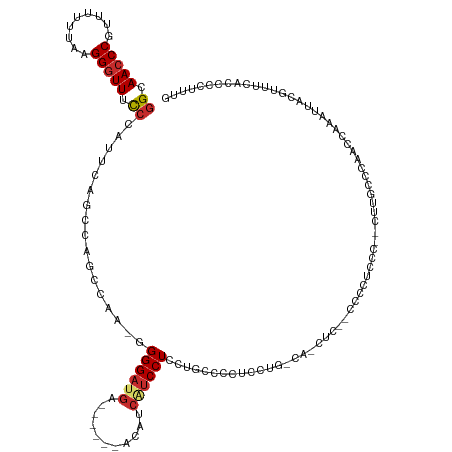
| Consensus MFE | -11.37 |
| Energy contribution | -11.87 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.15 |
| Mean z-score | -0.91 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.772944 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
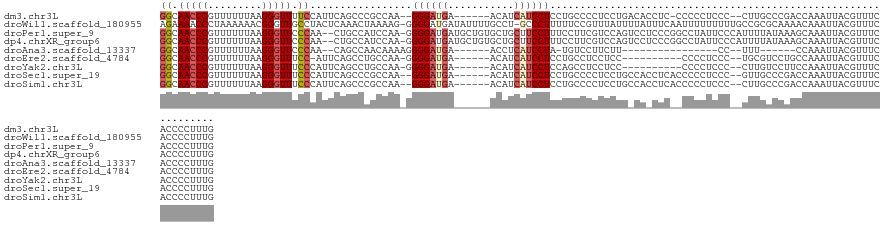
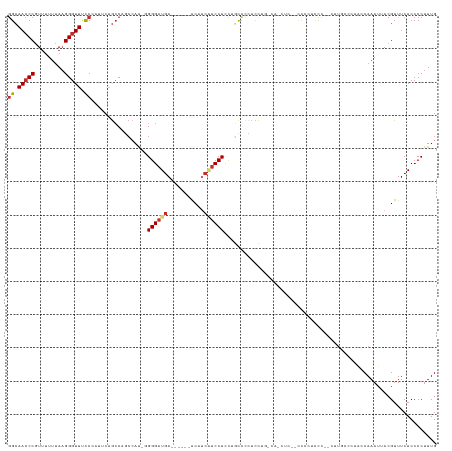
>dm3.chr3L 18845504 118 + 24543557 GGCAACCCGUUUUUUAAGGGUUUUCCAUUCAGCCCGCCAA--GGGAUGA------ACAUCAUCCUCCUGCCCCUCCUGACACCUC-CCCCCUCCC--CUUGCCCGACCAAAUUACGUUUCACCCCUUUG ((((((((.........)))).......((((...((...--((((((.------....))))))...)).....))))......-.........--..)))).......................... ( -21.70, z-score = -1.07, R) >droWil1.scaffold_180955 94155 127 + 2875958 AGAAAACCCUAAAAAACGGGUUGCCUACUCAAACUAAAAG-GGGGAUGAUAUUUUGCCU-GCCCUUUUUCCGUUUAUUUUAUUUCAAUUUUUUUUUGCCGCGCAAAACAAAUUACGUUUCACCCCUUUG ((..(((((........)))))..))..........((((-(((((((((.(((((((.-((..................................)).).))))))...))))....)).))))))). ( -21.58, z-score = 0.31, R) >droPer1.super_9 1235982 126 - 3637205 GGCAACCCGUUUUUUAAGGGUUCCCAA--CUGCCAUCCAA-GGGGAUGAUGCUGUGCUGCUUCCUUUCCUUCGUCCAGUCCUCCCGGCCUAUUCCCAUUUUAUAAAGCAAAUUACGUUUCACCCCUUUG ((((((((.........))))......--.))))...(((-((((.(((.((.(((.(((((..............((.((....)).))..............)))))...))))).))).))))))) ( -28.30, z-score = -0.90, R) >dp4.chrXR_group6 9202703 126 - 13314419 GGCAACCCGUUUUUUAAGGGUUCCCAA--CUGCCAUCCAA-GGGGAUGAUGCUGUGCUGCUUCCUUUCCUUCGUCCAGUCCUCCCGGCCUAUUCCCAUUUUAUAAAGCAAAUUACGUUUCACCCCUUUG ((((((((.........))))......--.))))...(((-((((.(((.((.(((.(((((..............((.((....)).))..............)))))...))))).))).))))))) ( -28.30, z-score = -0.90, R) >droAna3.scaffold_13337 4701142 96 - 23293914 GGCAACCCGUUUUUUAAGGGUUCCCAA--CAGCCAACAAAAGGGGAUGA------ACCUCAUCCUA-UGUCCUUCUU----------------CC--UUU------CCAAAUUACGUUUCACCCCUUUG ((.(((((.........))))).))..--.........(((((((.(((------(.(........-..........----------------..--...------.........).))))))))))). ( -20.05, z-score = -1.16, R) >droEre2.scaffold_4784 18662266 109 + 25762168 GGCAACCCGUUUUUUAAGGGUUUCC-AUUCAGCCUGCCAA-GGGGAUGA------ACAUCAUCCUCCUGCCUCCUCC----------CCCCUCCC--UGCGUCCUGCCAAAUUACGUUUCACCCCUUUG ((((...(((......((((.....-...((((((....)-))(((((.------....)))))..)))........----------.))))...--.)))...))))..................... ( -23.23, z-score = -0.71, R) >droYak2.chr3L 2398278 110 - 24197627 GGCAACCCGUUUUUUAAGGGUUUCCCAUUCAGCCUGCCAA-GGGGAUGA------ACAUCAUCCUCCAGCCUCCUCC----------CCCCUCCC--CUUGUCCUUCCAAAUUACGUUUCACCCCUUUG ((((((((.........)))))..................-(((((((.------....))))).)).)))......----------........--................................ ( -20.70, z-score = -0.64, R) >droSec1.super_19 811681 119 + 1257711 GGCAACCCGUUUUUUAAGGGUUUCCCAUUCAGCCCGCCAA--GGGAUGA------ACAUCAUCCUCCUGCCCCUCCUGCCACCUCACCCCCUCCC--GUUGCCCGACCAAAUUACGUUUCACCCCUUUG ((((((...........(((((........)))))((...--((((((.------....))))))...)).........................--)))))).......................... ( -24.60, z-score = -1.98, R) >droSim1.chr3L 18150284 119 + 22553184 GGCAACCCGUUUUUUAAGGGUUUCCCAUUCAGCCCGCCAA--GGGAUGA------ACAUCAUCCUCCUGCCCCUCCUGCCACCUCACCCCCUCCC--CUUGCCCGACCAAAUUACGUUUCACCCCUUUG ((((.............(((((........)))))((...--((((((.------....))))))...))......))))...............--................................ ( -21.70, z-score = -1.17, R) >consensus GGCAACCCGUUUUUUAAGGGUUUCCCAUUCAGCCAGCCAA_GGGGAUGA______ACAUCAUCCUCCUGCCCCUCCUG_CA_CUC__CCCCUCCC__CUUGCCCAACCAAAUUACGUUUCACCCCUUUG ((.(((((.........))))).)).................((((((...........))))))................................................................ (-11.37 = -11.87 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:36:31 2011