
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 18,641,734 – 18,641,893 |
| Length | 159 |
| Max. P | 0.625801 |

| Location | 18,641,734 – 18,641,854 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.65 |
| Shannon entropy | 0.16415 |
| G+C content | 0.40774 |
| Mean single sequence MFE | -27.65 |
| Consensus MFE | -20.47 |
| Energy contribution | -20.99 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.625801 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
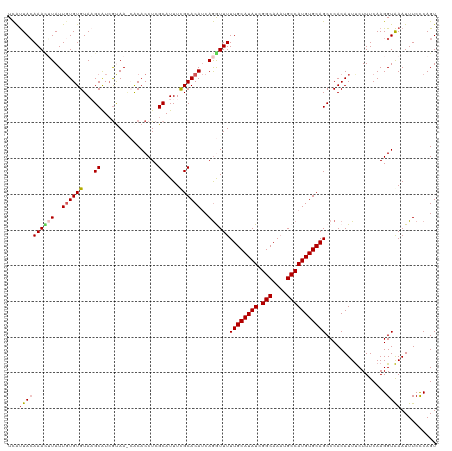
>dm3.chr3L 18641734 120 + 24543557 UCAUUACAAAUGUAUUGUGUGUAAGCAUCGUAAGCACACCUCGCAUUUCGCACAUCAGGUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCCCAGU ...((((((((((...((((((..((...))..))))))...)))))).((((((..(((.(((((......))))).)))))))))...................)))).......... ( -28.90, z-score = -2.27, R) >droEre2.scaffold_4784 10486905 118 - 25762168 UCGUUACAAAUUUAUUCUGUGUAUGCGUCGUAA--GCACCUCGCAUUUUGCACACCAGAUUUUUGCAUAUCGGCAAUUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCCCAGU .......(((((((((.((((((((((..((..--..))..)))))...)))))........((((((((.(((....)))))))))))................)))))))))...... ( -27.00, z-score = -2.04, R) >droYak2.chr3L 8779641 118 - 24197627 UUACUACAAAGGUAUUGUGUGUAUGCAUCAUAA--ACACCUCGCAUUUCGCACAUCAAAUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUCCUUUCAAGGUGAAUUUCCCAGU (((((..(((((...((((((.((((.......--.......))))..))))))........((((((((.(((....)))))))))))........)))))...))))).......... ( -29.44, z-score = -2.40, R) >droSec1.super_19 621872 120 + 1257711 UCAUUAAAAACUUAUUGUGUGUAAGCAUAGCAAACACACCUCGCAUUUUGCACAUCAGGUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCUCAGU .........((((...((((((..((...))..))))))........((((((((..(((.(((((......))))).)))))))))))...............))))............ ( -26.50, z-score = -1.65, R) >droSim1.chr3L 17952031 120 + 22553184 UCAUUACAAACUUAUUGUGUGUAAGCAUCGCAAACACACCUCGCAUUUCGCACAUCAGGUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCCCAGU ...((((.........((((((..((...))..))))))..........((((((..(((.(((((......))))).)))))))))...................)))).......... ( -26.40, z-score = -1.93, R) >consensus UCAUUACAAACUUAUUGUGUGUAAGCAUCGUAA_CACACCUCGCAUUUCGCACAUCAGGUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCCCAGU ...((((((((((..((((((...((................))....))))))..))))))((((((((.(((....))))))))))).................)))).......... (-20.47 = -20.99 + 0.52)


| Location | 18,641,774 – 18,641,893 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.23 |
| Shannon entropy | 0.35956 |
| G+C content | 0.38207 |
| Mean single sequence MFE | -25.12 |
| Consensus MFE | -14.42 |
| Energy contribution | -15.23 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.522979 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
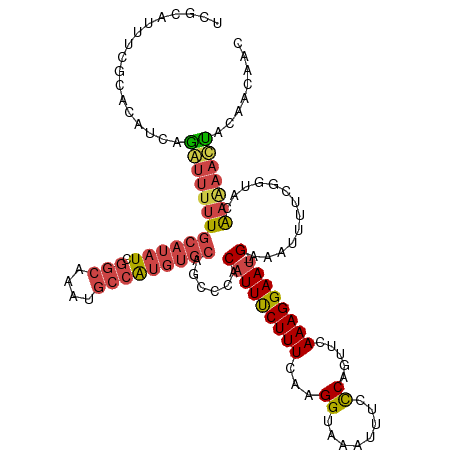
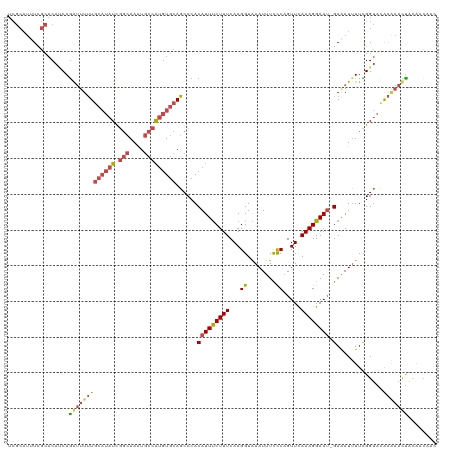
>dm3.chr3L 18641774 119 + 24543557 UCGCAUUUCGCACAUCAGGUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCCCAGUUCAAAGGAAU-GAAAUUUUCGGUACAAAAACUGCAACAAC ..(((....((((((..(((.(((((......))))).)))))))))......(((((((((...((........)).....))))))))-)...................)))...... ( -27.20, z-score = -1.83, R) >droAna3.scaffold_13337 4517240 105 - 23293914 -----AUUCUCUCUGCAGAAAUAU-----UUACCAACUUAAGCAUAUAUCCCACAUUUCUUUCAAGGUGAAUUUUCCUGUUUAAAGGAAAAGAAAUAUUUGG--CAGGCACCAUGAG--- -----......(((((..((((((-----((......................(((((......)))))..(((((((......))))))).)))))))).)--)))).........--- ( -18.80, z-score = -0.63, R) >droEre2.scaffold_4784 10486943 119 - 25762168 UCGCAUUUUGCACACCAGAUUUUUGCAUAUCGGCAAUUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCCCAGUUCAAAGGAAU-GAGAUUUUCGGUACGAUAAUUACAACAAC ..((.....))..(((......((((((((.(((....)))))))))))....(((((((((...((........)).....))))))))-)........)))................. ( -25.20, z-score = -1.50, R) >droYak2.chr3L 8779679 119 - 24197627 UCGCAUUUCGCACAUCAAAUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUCCUUUCAAGGUGAAUUUCCCAGUUCAAAGGAAU-GAAAUUUUUGGUACGAAAAUUACAACAAC .....(((((.((...((((((((((((((.(((....)))))))))))....((((((((......(((((......))))))))))))-)))))))...)).)))))........... ( -30.90, z-score = -3.06, R) >droSec1.super_19 621912 119 + 1257711 UCGCAUUUUGCACAUCAGGUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCUCAGUUCAAAGGAAU-GAAAUUUUCGGUAUAAAAACUACAACAAC ..((...((((((((..(((.(((((......))))).)))))))))))....(((((((((..(((......)))......))))))))-).........))................. ( -24.00, z-score = -1.54, R) >droSim1.chr3L 17952071 119 + 22553184 UCGCAUUUCGCACAUCAGGUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCCCAGUUCAAAGGAAU-GAAAUUUUCGGUACAAAAACUACAACAAC ..((.....))......(((((((((((((.(((....)))))))))......(((((((((...((........)).....))))))))-).............)))))))........ ( -24.60, z-score = -1.73, R) >consensus UCGCAUUUCGCACAUCAGAUUUUUGCAUAUCGGCAAAUGCCAUGUGCAGCCCACAUUUCUUUCAAGGUAAAUUUCCCAGUUCAAAGGAAU_GAAAUUUUCGGUACAAAAACUACAACAAC .................(((((((((((((.(((....))))))))).(((......(((....))).(((((((...((((....)))).)))))))..)))..)))))))........ (-14.42 = -15.23 + 0.81)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:36:09 2011