| Sequence ID | dm3.chr3L |
|---|---|
| Location | 18,641,220 – 18,641,300 |
| Length | 80 |
| Max. P | 0.952565 |

| Location | 18,641,220 – 18,641,300 |
|---|---|
| Length | 80 |
| Sequences | 10 |
| Columns | 80 |
| Reading direction | forward |
| Mean pairwise identity | 80.49 |
| Shannon entropy | 0.42380 |
| G+C content | 0.40267 |
| Mean single sequence MFE | -14.26 |
| Consensus MFE | -10.28 |
| Energy contribution | -10.04 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.952565 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
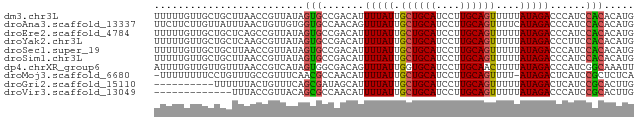
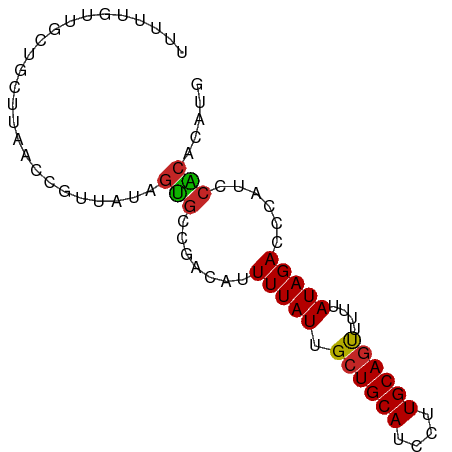
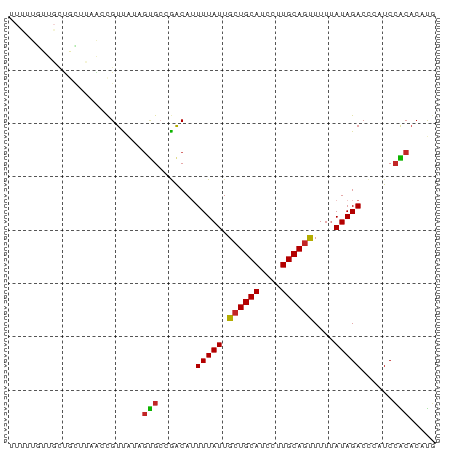
>dm3.chr3L 18641220 80 + 24543557 UUUUUGUUGCUGCUUAACCGUUAUAGUGCCGACAUUUUAUUGCUGCAUCCUUGCAGUUUUUAUAGACCCAUCCACACAUG ....(((((..(((..........)))..))))).(((((.((((((....))))))....))))).............. ( -13.80, z-score = -1.57, R) >droAna3.scaffold_13337 4516678 80 - 23293914 UUCUUCUUGUUAUUUAACUGUUGUGGUGCCAACAGUUUAUUGCUGCAUCCUUGCAGUUUUCAUAGACCCAUCCACACAUG ..................(((.((((........((((((.((((((....))))))....))))))....))))))).. ( -16.60, z-score = -1.76, R) >droEre2.scaffold_4784 10486320 80 - 25762168 UUUUUGUUGCUGCUCAGCCGUUAUAGUGCCGACAUUUUAUUGCUGCAUCCUUGCAGUUUUUAUAGACCCAUCCACACAUG ........(((....))).......(((..((...(((((.((((((....))))))....)))))....))..)))... ( -13.80, z-score = -0.87, R) >droYak2.chr3L 8779053 80 - 24197627 UUUUUGUUGCUGCUCAAGCGUUAUAGUGCCGACAUUUUAUUGCUGCAUCCUUGCAGUUUUUAUAGACCCUUCCACACAUG ........(((.....)))......(((..((...(((((.((((((....))))))....)))))....))..)))... ( -14.90, z-score = -1.20, R) >droSec1.super_19 621328 80 + 1257711 UUUUUGUUGCUGCUUAACCGUUAUAGUGCCGACAUUUUAUUGCUGCAUCCUUGCAGUUUUUAUAGACCCAUCCACACAUG ....(((((..(((..........)))..))))).(((((.((((((....))))))....))))).............. ( -13.80, z-score = -1.57, R) >droSim1.chr3L 17951503 80 + 22553184 UUUUUGUUGCUGCUUAACCGUUAUAGUGCCGACAUUUUAUUGCUGCAUCCUUGCAGUUUUUAUAGACCCAUCCACACAUG ....(((((..(((..........)))..))))).(((((.((((((....))))))....))))).............. ( -13.80, z-score = -1.57, R) >dp4.chrXR_group6 11531488 80 + 13314419 AUUUUGUUGUUGUUUAACCGUCAUAGUGGCGACAGUUUAUUGGUGCAUCCUUGCAACUUUUAUAGACCCAUCGGCAAAUU ..(((((((.((......((((.....))))...((((((.((((((....))).)))...)))))).)).))))))).. ( -18.40, z-score = -1.34, R) >droMoj3.scaffold_6680 16685338 78 - 24764193 -UUUUUUUUCCUGUUUGCCGUUUCAACGCCAACAUUUUAUUGCUGCAUCCUUGCAGUUUU-AUAGACUCAUCCGCUCUCA -..........((((.((.((....)))).)))).(((((.((((((....))))))...-))))).............. ( -12.70, z-score = -2.53, R) >droGri2.scaffold_15110 16529427 70 + 24565398 ----------UUUUUUACUGUUUCAGCGAUAGCAUUUUAUUGCUGCAUCCUUGCAGUUUUUAUAGACUCAUCCGCACUUG ----------...............(((.......(((((.((((((....))))))....)))))......)))..... ( -12.42, z-score = -1.62, R) >droVir3.scaffold_13049 4862869 67 - 25233164 -------------UUUACCGUUACAGCGCCAACAUUUUAUUGCUGCAUCCUUGCAGUUUUUAUAGACCCAUCCGCACUUG -------------............(((.......(((((.((((((....))))))....)))))......)))..... ( -12.42, z-score = -2.94, R) >consensus UUUUUGUUGCUGCUUAACCGUUAUAGUGCCGACAUUUUAUUGCUGCAUCCUUGCAGUUUUUAUAGACCCAUCCACACAUG .........................(((.......(((((.((((((....))))))....)))))......)))..... (-10.28 = -10.04 + -0.24)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:36:07 2011