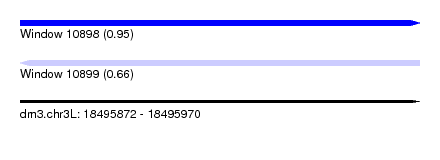

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 18,495,872 – 18,495,970 |
| Length | 98 |
| Max. P | 0.947538 |

| Location | 18,495,872 – 18,495,970 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 84.15 |
| Shannon entropy | 0.28397 |
| G+C content | 0.47538 |
| Mean single sequence MFE | -33.56 |
| Consensus MFE | -26.34 |
| Energy contribution | -27.22 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.947538 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
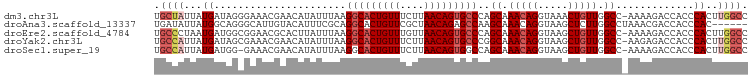
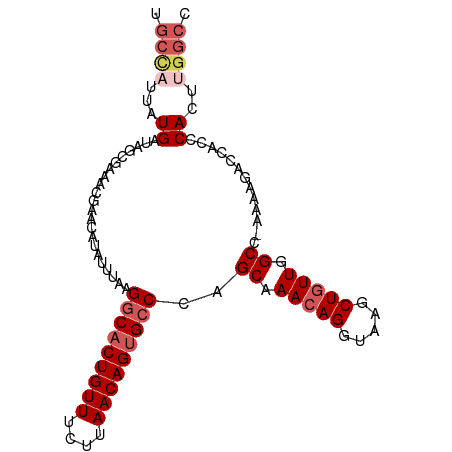
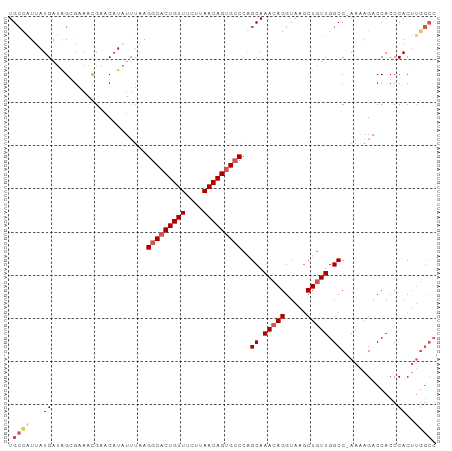
>dm3.chr3L 18495872 98 + 24543557 GGCCAAGUGGGUGGUCUUUU-GGCCAACAGUUUACCUGUUUGCUGGGCACUGUUAAGAAACAGUGCCUUAAAUAUGUUCGUUUCCCUAUCAUAAUAGCA (((((((.((....)).)))-))))(((((.....)))))((((((((((((((....))))))))).....((((.............)))).))))) ( -31.52, z-score = -2.13, R) >droAna3.scaffold_13337 4394708 93 - 23293914 ------GUGGGUGGUCGUUUAGGCCAAGAGCUUACCUGUUUGCUUGGCUCUGUUAGCGAACAGUGCCUGCGAAAUGUACAAUGCCCUGCCAUAAUAUCA ------((((.(((((.....)))))((.((.(((...(((((..(((.(((((....))))).))).)))))..)))....)).)).))))....... ( -29.50, z-score = -1.06, R) >droEre2.scaffold_4784 10361848 98 - 25762168 GGCCAAGUGGGUGGUCUUUU-GGCCAACAGCUUACCUGUUUGCUGGGCACUGUUAACAAACAGUGCCUUAAAUAAGUGCGUUCCGCCAUCAUUAGGGCA .(((.....((((((.....-(((.(((((.....))))).)))((((((((((....))))))))))................)))))).....))). ( -36.50, z-score = -1.82, R) >droYak2.chr3L 8650520 98 - 24197627 GGCCAAGUGGGUGGUCUCUU-GGCCAACAGCUUACCUGUUUGCCGGGCACUGUUAAGAAACAGUGCCUUAAAUAUGUUCGUUUCGCUAUCAUAAUGGCA .((((....((((((.....-(((.(((((.....))))).)))((((((((((....))))))))))................))))))....)))). ( -37.30, z-score = -3.35, R) >droSec1.super_19 496775 97 + 1257711 GGCCAAGUGGGUGGUCUUUU-GGCCAACAGCUUACCUGUUUGCUGGCCACUGUUAAGAAACAGUGCCUUAAAUAUGUUCGUUUC-CCAUCAUAAUGGCA .((((.((((((((((....-)))))..(((..((.((((((..(((.((((((....))))))))).)))))).))..))).)-)))).....)))). ( -33.00, z-score = -2.21, R) >consensus GGCCAAGUGGGUGGUCUUUU_GGCCAACAGCUUACCUGUUUGCUGGGCACUGUUAAGAAACAGUGCCUUAAAUAUGUUCGUUUCCCUAUCAUAAUGGCA .(((.....(((((.......(((.(((((.....))))).)))((((((((((....)))))))))).................))))).....))). (-26.34 = -27.22 + 0.88)



| Location | 18,495,872 – 18,495,970 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 84.15 |
| Shannon entropy | 0.28397 |
| G+C content | 0.47538 |
| Mean single sequence MFE | -29.66 |
| Consensus MFE | -18.96 |
| Energy contribution | -20.44 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.664560 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 18495872 98 - 24543557 UGCUAUUAUGAUAGGGAAACGAACAUAUUUAAGGCACUGUUUCUUAACAGUGCCCAGCAAACAGGUAAACUGUUGGCC-AAAAGACCACCCACUUGGCC ((((..((((.(..(....).).)))).....(((((((((....))))))))).))))(((((.....)))))((((-((..(......)..)))))) ( -32.00, z-score = -3.41, R) >droAna3.scaffold_13337 4394708 93 + 23293914 UGAUAUUAUGGCAGGGCAUUGUACAUUUCGCAGGCACUGUUCGCUAACAGAGCCAAGCAAACAGGUAAGCUCUUGGCCUAAACGACCACCCAC------ .........((((((((....(((.....((.(((.(((((....))))).)))..))......))).)))))..)))...............------ ( -21.30, z-score = 0.36, R) >droEre2.scaffold_4784 10361848 98 + 25762168 UGCCCUAAUGAUGGCGGAACGCACUUAUUUAAGGCACUGUUUGUUAACAGUGCCCAGCAAACAGGUAAGCUGUUGGCC-AAAAGACCACCCACUUGGCC (((...(((((..(((...)))..)))))...(((((((((....)))))))))..)))(((((.....)))))((((-((..(......)..)))))) ( -30.00, z-score = -0.90, R) >droYak2.chr3L 8650520 98 + 24197627 UGCCAUUAUGAUAGCGAAACGAACAUAUUUAAGGCACUGUUUCUUAACAGUGCCCGGCAAACAGGUAAGCUGUUGGCC-AAGAGACCACCCACUUGGCC ((((..((((....((...))..)))).....(((((((((....))))))))).))))(((((.....)))))((((-(((.(......).))))))) ( -35.00, z-score = -4.05, R) >droSec1.super_19 496775 97 - 1257711 UGCCAUUAUGAUGG-GAAACGAACAUAUUUAAGGCACUGUUUCUUAACAGUGGCCAGCAAACAGGUAAGCUGUUGGCC-AAAAGACCACCCACUUGGCC .((((.((((.(.(-....).).)))).....(((((((((....))))))((((((((...........))))))))-....).)).......)))). ( -30.00, z-score = -1.89, R) >consensus UGCCAUUAUGAUAGCGAAACGAACAUAUUUAAGGCACUGUUUCUUAACAGUGCCCAGCAAACAGGUAAGCUGUUGGCC_AAAAGACCACCCACUUGGCC .((((...((......................(((((((((....)))))))))..((.(((((.....))))).)).............))..)))). (-18.96 = -20.44 + 1.48)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:35:58 2011