
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 18,383,627 – 18,383,723 |
| Length | 96 |
| Max. P | 0.693553 |

| Location | 18,383,627 – 18,383,723 |
|---|---|
| Length | 96 |
| Sequences | 11 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 80.68 |
| Shannon entropy | 0.41572 |
| G+C content | 0.46287 |
| Mean single sequence MFE | -22.94 |
| Consensus MFE | -14.85 |
| Energy contribution | -15.15 |
| Covariance contribution | 0.30 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.693553 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 18383627 96 - 24543557 -CGGAUCUUCCAAGGAGGCUCAGU---CAGGCAGUAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUGCCCCAACCCCAA-CCUCAUU -.((.....))...((((....((---..((((((......))((((((((((((......)))))).)))))).....))))...)).....-))))... ( -23.10, z-score = -1.28, R) >droSim1.chr3L 17718747 96 - 22553184 -CGGAUCUUCCAAGGAGGCUCAGU---CAGGCAGUAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUGCCCCAACC-CAAACCUCAUU -.((.....))...((((....((---..((((((......))((((((((((((......)))))).)))))).....))))...)).-....))))... ( -23.10, z-score = -1.27, R) >droSec1.super_19 379245 96 - 1257711 -CGGAUCUUCCAAGGAGGCUCAGU---CAGGCAGUAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUGCCCCAACC-CAAACCUCAUU -.((.....))...((((....((---..((((((......))((((((((((((......)))))).)))))).....))))...)).-....))))... ( -23.10, z-score = -1.27, R) >droYak2.chr3L 8538388 96 + 24197627 -CGGAUCUUCCAAGGAGGCUCAGU---CAGGCAGUAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUGCCCCAAAC-CAAACCUCAUU -.((.....))...((((....((---..((..(((((.(...((((((((((((......)))))).))))))..)))))).))..))-....))))... ( -23.10, z-score = -1.36, R) >droEre2.scaffold_4784 10253474 96 + 25762168 -CGGAUCUUCCAAGGAGGCUCAGU---CAGGCAGUAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUGCCCCAACC-CAAACCUCAUU -.((.....))...((((....((---..((((((......))((((((((((((......)))))).)))))).....))))...)).-....))))... ( -23.10, z-score = -1.27, R) >droAna3.scaffold_13337 4283160 97 + 23293914 -AGGAUCUUGCAAGGAGGCUCAGU---CAGGCAGUAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUGCCCCAACCACAAGCCUUAUU -.............((((((..((---..((((((......))((((((((((((......)))))).)))))).....))))...))....))))))... ( -23.80, z-score = -0.81, R) >dp4.chrXR_group6 11277468 97 - 13314419 AAGGACACUCGUAGGAGGCUCAGC---CAGGCGGCAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUACCUCAGUCCCAAUCCCAGU- ..((((.(.....)((((....((---(....)))........((((((((((((......)))))).)))))).......)))).))))..........- ( -26.90, z-score = -1.60, R) >droPer1.super_20 1178084 97 - 1872136 AAGGACACUCGUAGGAGGCUCAGC---CAGGCGGCAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUACCUCAGUCCCAGUCCCAGU- ..((((........((((....((---(....)))........((((((((((((......)))))).)))))).......)))).......))))....- ( -27.66, z-score = -1.52, R) >droWil1.scaffold_180949 5876314 87 + 6375548 ------------AGGAUUUUCGUCUUUCAAGCAGCAAAUGAACCACUAAUUGCGUGCCGAAAUGCAAUUUAGUGAACUUUACCU-AGUCCGAGCGCCCUC- ------------.((((((((((..............))))).((((((((((((......)))))).))))))..........-)))))..........- ( -16.94, z-score = -0.36, R) >droVir3.scaffold_13049 21578196 85 - 25233164 ---------CGAAGCCAGCUCAGC---CACGCAGCAAAUGAACCACUAAUUGCGUGCCGAAAUGCAAUUUAGUGAACUUUGCCCCAGACCCAAAAAC---- ---------(((((...((((...---...).)))........((((((((((((......)))))).))))))..)))))................---- ( -17.00, z-score = -1.57, R) >droGri2.scaffold_15110 2541046 92 + 24565398 -UAUGUAUUUGUGGGAGGCUCAGC---CACGCAGCAAAUGAACCACUAAUUGCGUGCCGAAAUGCAAUUUAGUGAACUUUACCCCAAACCCAAAAU----- -..((..((((.(((.(((...))---)...............((((((((((((......)))))).)))))).......)))))))..))....----- ( -24.50, z-score = -2.32, R) >consensus _CGGAUCUUCCAAGGAGGCUCAGU___CAGGCAGUAAAUGAACCACUAAUUGCGUGACGAAAUGCAAUUUAGUGAACUUUGCCCCAACCCCAAACCUCAUU .............((.(((..((........((.....))...((((((((((((......)))))).))))))..))..)))))................ (-14.85 = -15.15 + 0.30)

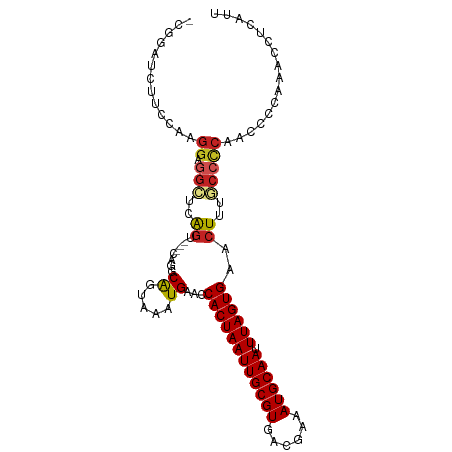

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:35:43 2011