
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 18,073,669 – 18,073,801 |
| Length | 132 |
| Max. P | 0.873923 |

| Location | 18,073,669 – 18,073,801 |
|---|---|
| Length | 132 |
| Sequences | 6 |
| Columns | 138 |
| Reading direction | forward |
| Mean pairwise identity | 77.13 |
| Shannon entropy | 0.42622 |
| G+C content | 0.36680 |
| Mean single sequence MFE | -30.22 |
| Consensus MFE | -19.31 |
| Energy contribution | -18.65 |
| Covariance contribution | -0.65 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.873923 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 18073669 132 + 24543557 CUUUAUAUUUCCCCCACUUUACAAUAGGUUAGUCUGUUAUUGUUCUGUCUGUGAGGACUACAAUUUGUCCGUGUGCGCUUGUUUAUAUAUGUAUGU----AUGUAUCUCCGCAUAUAUAGGAAAGUACAGAAUAUG-- ......................((((((....))))))..((((((((((....((((........))))(((((((......(((((((....))----)))))....))))))).......)).))))))))..-- ( -28.70, z-score = -1.03, R) >droSim1.chr3L 17400305 124 + 22553184 CUUUAUAUUUCCCCCACUUUACAAUAGGUUAGUCUGUUAUUGUUCUGUCUGUGAGGACUACAAUUUGUCCGUGUGCGCUUGUUUAUAUAUGUAU------------CUCCGCAUAUAUAGGAAAGUACAGAAUAUG-- ...(((((((....(((.(((((((((..((((.....))))..)))).)))))((((........)))))))((..(((.(((((((((((..------------....))))))))))).)))..)))))))))-- ( -29.60, z-score = -2.19, R) >droSec1.super_19 69843 124 + 1257711 CUUUGUAUUUCCCCCACUUUACAAUAGGUUAGUCUGUUAUUGUUCUGUCUGUGAGGACUACAAUUUGUCCGUGUGCGCUUGUUUAUAUAUGUAU------------CUCCGCAUAUAUAGGAAAGUACAGAAUAUG-- .(((((((((.((.(((.(((((((((..((((.....))))..)))).)))))((((........)))))))..........(((((((((..------------....))))))))))).))))))))).....-- ( -30.90, z-score = -2.47, R) >droYak2.chr3L 8214653 135 - 24197627 CUUUUUGUUUCCCCCACUUUACAAUAGGUUAGUCUGUUAUUGUUCUGUCUGCGAGGAUUAAAAUUCGUCCAUGUGCGCUUGUUUUUAUAUAUAUCUCCGUAUA---UGUAUAAUAUAUGGGAGAGCACAUGGCGUAUG ..(((((..(((.(((..(.(((((((.........)))))))...)..)).).))).)))))..((.((((((((.((..(..((((((((((......)))---)))))))...)..))...)))))))))).... ( -32.20, z-score = -1.71, R) >droEre2.scaffold_4784 9945028 138 - 25762168 CUUUAUGUUUCCCCCACUUUACAAUAGGUUAGUCUGUUAUUGUUCUGUCUGUGAGGACUCAAAUUUGUCCGUGUGCGUUUGUUUAUACUCGUAUUUGUAUAUGCAUCUCCGCAUAUAUGGGAGAGUACAAAGCGUAUG .....((((((.((((..(((((((((..((((.....))))..)))).)))))((((........))))(((((((..(((.(((((........))))).)))....))))))).)))).))).)))......... ( -35.70, z-score = -2.47, R) >droAna3.scaffold_13337 18135216 126 - 23293914 CUUUAUAUGUCCCCCACUUUACAAUAGGUUAGUCUGUUAUUGUUCUGUCUGUGUGGAUUAAAAUAUAGAGGUAGAGAUGUGUGAGUA-GUAUCUUC--------AUCUAUGUAUGUAUAUGUAUCUCCAC-GUAUG-- ((((((((.....((((...(((((((..((((.....))))..)))).)))))))......))))))))((.((((((..((....-((((.(..--------....).))))...))..)))))).))-.....-- ( -24.20, z-score = 0.20, R) >consensus CUUUAUAUUUCCCCCACUUUACAAUAGGUUAGUCUGUUAUUGUUCUGUCUGUGAGGACUAAAAUUUGUCCGUGUGCGCUUGUUUAUAUAUGUAU_U__________CUCCGCAUAUAUAGGAAAGUACAGAAUAUG__ ...(((((((......(((((((((((..((((.....))))..)))).)))))))...............(((((.....(((((((((((..................)))))))))))...)))))))))))).. (-19.31 = -18.65 + -0.65)

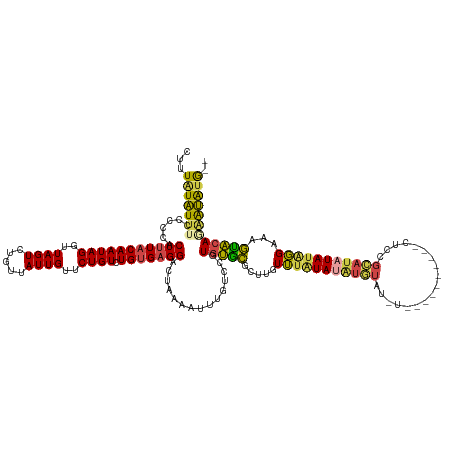
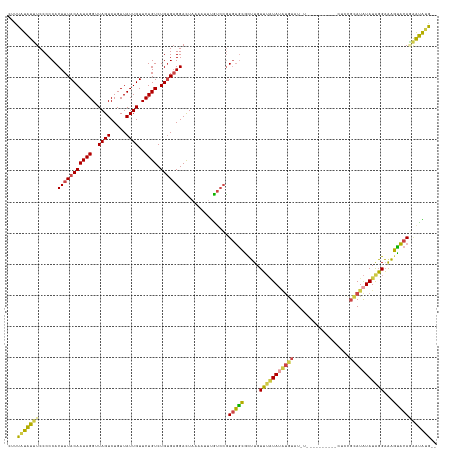
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:35:08 2011