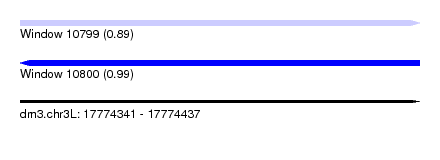

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 17,774,341 – 17,774,437 |
| Length | 96 |
| Max. P | 0.993389 |

| Location | 17,774,341 – 17,774,437 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 80.00 |
| Shannon entropy | 0.36412 |
| G+C content | 0.51908 |
| Mean single sequence MFE | -32.74 |
| Consensus MFE | -25.62 |
| Energy contribution | -25.44 |
| Covariance contribution | -0.18 |
| Combinations/Pair | 1.32 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.887972 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
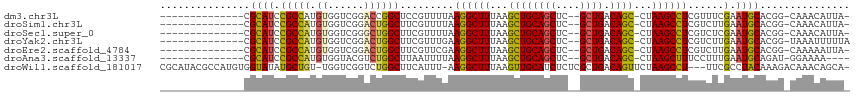
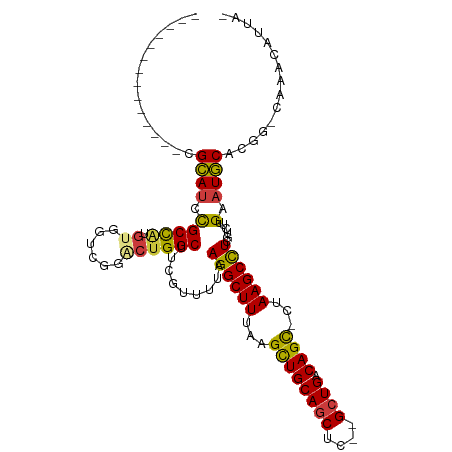
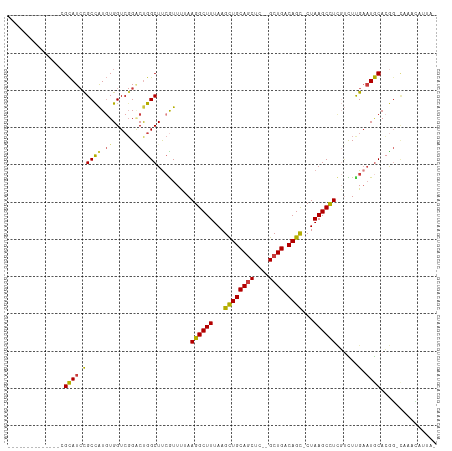
>dm3.chr3L 17774341 96 + 24543557 --------------CGCAUCCGCCAUGUGGUCGGACCGGCUCCGUUUUAAGGCUUUAAGCUGCAGCUC--GCUGACAGC-CUAAGCCUCGUUUCGAAUGCACGG-CAAACAUUA- --------------....((((((....)).))))(((((..((.....((((((...(((((((...--.))).))))-..)))))).....))...)).)))-.........- ( -33.00, z-score = -1.44, R) >droSim1.chr3L 17111891 96 + 22553184 --------------CGCAUCCGCCAUGUGGUCGGACUGGCUUCGUUUUAAGGCUUUAAGCUGCAGCUC--GCUGACAGC-CUAAGCCUCGUCUUGAAUGCACGG-CAAACAUUA- --------------.((.((((((....)).))))...((((((.....((((((...(((((((...--.))).))))-..)))))).....)))).))...)-)........- ( -32.60, z-score = -1.35, R) >droSec1.super_0 9821392 96 + 21120651 --------------CGCAUCCGCCAUGUGGUCGGGCUGGCUUCGUUUUAAGGCUUUAAGCUGCAGCUC--GCUGACAGC-CUAAGCCUCGUCUCGAAUGCACGG-CAAACAUUA- --------------.....(((((....)).)))((((((((((.....((((((...(((((((...--.))).))))-..)))))).....)))).)).)))-)........- ( -36.90, z-score = -2.26, R) >droYak2.chr3L 7914998 97 - 24197627 --------------CGCAUCCGCCAUGUGGUCGGACUGGCUUCGUUUGAAGGCUUUAAGCUGCAGCUC--GCUGACAGC-CUAAGCCUCGUCUUGAAUGCACGG-UAAAUUUUUA --------------.....(((((....)).)))((((((((((..(((.(((((...(((((((...--.))).))))-..))))))))...)))).)).)))-)......... ( -33.60, z-score = -1.72, R) >droEre2.scaffold_4784 9675926 96 - 25762168 --------------CGCAUCCGCCAUGUGGUCGGACUGGCUUCGUUCGAAGGCUUUAAGCUGCAGCUC--GCUGACAGC-CUAAGCCUCGUCUUGAAUGCACGG-CAAAAAUUA- --------------.((.((((((....)).))))...((((((..(((.(((((...(((((((...--.))).))))-..))))))))...)))).))...)-)........- ( -35.20, z-score = -1.93, R) >droAna3.scaffold_13337 14471435 93 + 23293914 --------------CGCAUCCGCCAUGUGGUACGUCUGGCUUAAUUUUAAGGCUUUAAGCUGCAGCUC--GCUGACAGC-CUAAGCUUUCCUUUGAAUGCAGAU-GGAAAA---- --------------....((((((....)))..(((((..((((....(((((((...(((((((...--.))).))))-..)))))))...))))...)))))-)))...---- ( -27.10, z-score = -0.65, R) >droWil1.scaffold_181017 696765 109 - 939937 CGCAUACGCCAUGUGGUAUAUGCUGU-UGGUCGGUCUGGCUUCAUUU-AAGGCUUUAAGUUGCAUCUCUCGCUGACAGUUCUAAGCCU---UUCGCCUACAAAGACAAACAGCA- ...(((..((....))..)))(((((-(.(((.((..(((.......-(((((((..(..((((........)).))..)..))))))---)..))).))...))).)))))).- ( -30.80, z-score = -1.22, R) >consensus ______________CGCAUCCGCCAUGUGGUCGGACUGGCUUCGUUUUAAGGCUUUAAGCUGCAGCUC__GCUGACAGC_CUAAGCCUCGUCUUGAAUGCACGG_CAAACAUUA_ ...............((((.(((((.((......)))))).........((((((...((((((((....)))).))))...))))))......).))))............... (-25.62 = -25.44 + -0.18)
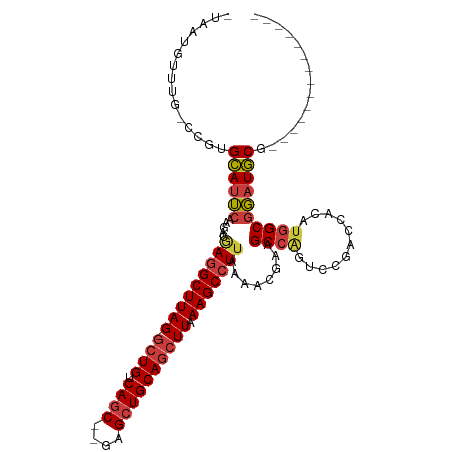
| Location | 17,774,341 – 17,774,437 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 80.00 |
| Shannon entropy | 0.36412 |
| G+C content | 0.51908 |
| Mean single sequence MFE | -34.87 |
| Consensus MFE | -28.41 |
| Energy contribution | -29.00 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.61 |
| SVM RNA-class probability | 0.993389 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
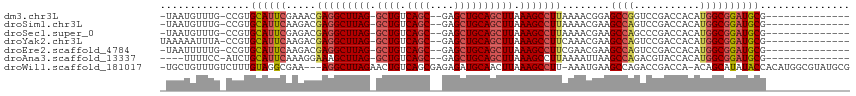
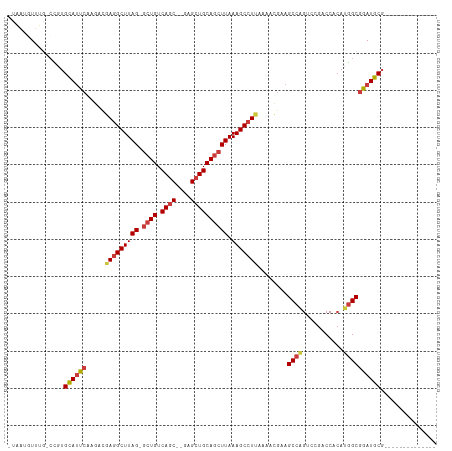
>dm3.chr3L 17774341 96 - 24543557 -UAAUGUUUG-CCGUGCAUUCGAAACGAGGCUUAG-GCUGUCAGC--GAGCUGCAGCUUAAAGCCUUAAAACGGAGCCGGUCCGACCACAUGGCGGAUGCG-------------- -........(-(((.((.((((....(((((((((-((((.(((.--...))))))))).)))))))....))))))))(((((.((....))))))))).-------------- ( -39.40, z-score = -2.82, R) >droSim1.chr3L 17111891 96 - 22553184 -UAAUGUUUG-CCGUGCAUUCAAGACGAGGCUUAG-GCUGUCAGC--GAGCUGCAGCUUAAAGCCUUAAAACGAAGCCAGUCCGACCACAUGGCGGAUGCG-------------- -.........-....((((((.....(((((((((-((((.(((.--...))))))))).)))))))........((((((......)).)))))))))).-------------- ( -36.40, z-score = -2.63, R) >droSec1.super_0 9821392 96 - 21120651 -UAAUGUUUG-CCGUGCAUUCGAGACGAGGCUUAG-GCUGUCAGC--GAGCUGCAGCUUAAAGCCUUAAAACGAAGCCAGCCCGACCACAUGGCGGAUGCG-------------- -.........-....((((((.....(((((((((-((((.(((.--...))))))))).)))))))........((((...........)))))))))).-------------- ( -35.70, z-score = -2.14, R) >droYak2.chr3L 7914998 97 + 24197627 UAAAAAUUUA-CCGUGCAUUCAAGACGAGGCUUAG-GCUGUCAGC--GAGCUGCAGCUUAAAGCCUUCAAACGAAGCCAGUCCGACCACAUGGCGGAUGCG-------------- ..........-....((((((.....(((((((((-((((.(((.--...))))))))).)))))))........((((((......)).)))))))))).-------------- ( -35.80, z-score = -3.33, R) >droEre2.scaffold_4784 9675926 96 + 25762168 -UAAUUUUUG-CCGUGCAUUCAAGACGAGGCUUAG-GCUGUCAGC--GAGCUGCAGCUUAAAGCCUUCGAACGAAGCCAGUCCGACCACAUGGCGGAUGCG-------------- -.........-....((((((....((((((((((-((((.(((.--...))))))))).))))).)))......((((((......)).)))))))))).-------------- ( -37.20, z-score = -2.74, R) >droAna3.scaffold_13337 14471435 93 - 23293914 ----UUUUCC-AUCUGCAUUCAAAGGAAAGCUUAG-GCUGUCAGC--GAGCUGCAGCUUAAAGCCUUAAAAUUAAGCCAGACGUACCACAUGGCGGAUGCG-------------- ----......-....((((((.((((.....((((-((((.(((.--...)))))))))))..))))........((((...........)))))))))).-------------- ( -30.30, z-score = -2.28, R) >droWil1.scaffold_181017 696765 109 + 939937 -UGCUGUUUGUCUUUGUAGGCGAA---AGGCUUAGAACUGUCAGCGAGAGAUGCAACUUAAAGCCUU-AAAUGAAGCCAGACCGACCA-ACAGCAUAUACCACAUGGCGUAUGCG -...((((.(((...((.(((..(---((((((.((..(((...(....)..)))..)).)))))))-.......)))..)).))).)-)))(((((..((....))..))))). ( -29.30, z-score = -1.13, R) >consensus _UAAUGUUUG_CCGUGCAUUCAAGACGAGGCUUAG_GCUGUCAGC__GAGCUGCAGCUUAAAGCCUUAAAACGAAGCCAGUCCGACCACAUGGCGGAUGCG______________ ...............((((((.....(((((((...((((.((((....))))))))...)))))))........((((...........))))))))))............... (-28.41 = -29.00 + 0.59)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:34:36 2011