
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 17,340,270 – 17,340,357 |
| Length | 87 |
| Max. P | 0.608411 |

| Location | 17,340,270 – 17,340,357 |
|---|---|
| Length | 87 |
| Sequences | 7 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 66.86 |
| Shannon entropy | 0.63333 |
| G+C content | 0.41875 |
| Mean single sequence MFE | -21.31 |
| Consensus MFE | -7.37 |
| Energy contribution | -9.47 |
| Covariance contribution | 2.10 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.608411 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
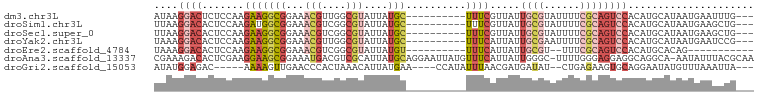
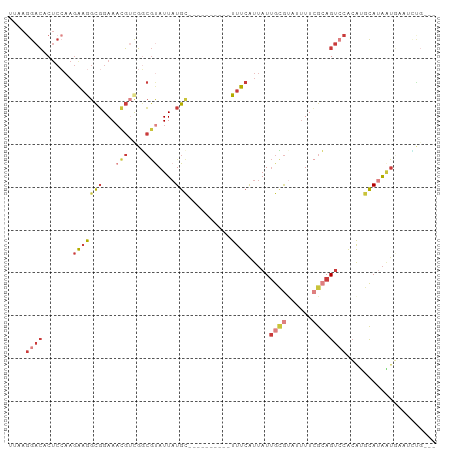
>dm3.chr3L 17340270 87 + 24543557 AUAAGGACUCUCCAAGAAGGCGGAAACGUUGGCGUAUUAUGC----------UUUCGUUAUUGCGUAUUUUCGCAGUCCACAUGCAUAAUGAAUUUG--- ....((.....))..(..((((....))))..).((((((((----------.......((((((......))))))......))))))))......--- ( -23.22, z-score = -1.72, R) >droSim1.chr3L 16680484 87 + 22553184 UUAAGGACACUCCAAGAUGGCGGAAACGUCGGCGUAUUAUGC----------UUUCGUUAUUGCGUAUUUUCGCAGUCCACAUGCAUAAUGAAGCUG--- ....((.....)).....((((....))))(((.((((((((----------.......((((((......))))))......))))))))..))).--- ( -26.72, z-score = -2.35, R) >droSec1.super_0 9391601 87 + 21120651 UUAAGGACACUCCAAGAAGGCGGAAACGUCGGCGUAUUAUGC----------UUUCGUUAUUGCGUAUUUUCGCAGUCCACAUGCAUAAUGAAGCUG--- ....((.....)).....((((....))))(((.((((((((----------.......((((((......))))))......))))))))..))).--- ( -26.92, z-score = -2.56, R) >droYak2.chr3L 7468606 87 - 24197627 UAAAGGACACUCCAAGAAGGCGGAAACGUUGGCGUAUUAUGC----------UUUCAUUAUUGCGAAUUUUCGCAGUCCACAUGCAUAAUGAAUCCG--- ....(((........(..((((....))))..).((((((((----------.......(((((((....)))))))......))))))))..))).--- ( -25.82, z-score = -2.98, R) >droEre2.scaffold_4784 9253325 77 - 25762168 UAAAGGACACUCCAAGAAGGCGGAAACGUCGGCGUAUUAUGU----------UUUCAUUAUUGCGU--UUUCGCAGUCCACAUGCACAG----------- ....((((..........((((....))))((((((..(((.----------...)))...)))))--)......))))..........----------- ( -20.80, z-score = -2.34, R) >droAna3.scaffold_13337 8294455 98 + 23293914 CGAAAGACACUCGAAGGAAGCGGAAAUGACGUCGCAUUAUGCAGGAAUUAUGUUUCAUUAUUGGGC-UUUUGGGAGGAGGCAGGCA-AAUAUUUACGCAA (....)....((....)).(((.(((((..(((((.....)).(((((...)))))........((-((((....)))))).))).-..))))).))).. ( -17.60, z-score = 0.30, R) >droGri2.scaffold_15053 219494 86 - 344661 AUAUGGAGAC-----AAAAGUUGAACCCACUAAACAUUAUGAA----CCAUAUUUAACGAUGAUAU--CUGAGAAGUGCAGGAAUAUGUUUAAAUUA--- ...((....)-----).......................((((----(.((((((.....((....--((....))..)).))))))))))).....--- ( -8.10, z-score = 0.93, R) >consensus UUAAGGACACUCCAAGAAGGCGGAAACGUCGGCGUAUUAUGC__________UUUCAUUAUUGCGUAUUUUCGCAGUCCACAUGCAUAAUGAAUCUG___ ....((((..........((((....))))...............................((((......))))))))..................... ( -7.37 = -9.47 + 2.10)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:33:32 2011