
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 5,519,909 – 5,519,987 |
| Length | 78 |
| Max. P | 0.845050 |

| Location | 5,519,909 – 5,519,987 |
|---|---|
| Length | 78 |
| Sequences | 5 |
| Columns | 78 |
| Reading direction | forward |
| Mean pairwise identity | 72.68 |
| Shannon entropy | 0.49664 |
| G+C content | 0.40642 |
| Mean single sequence MFE | -16.63 |
| Consensus MFE | -9.48 |
| Energy contribution | -10.84 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.845050 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
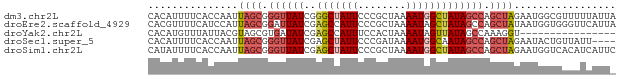
>dm3.chr2L 5519909 78 + 23011544 CACAUUUUCACCAAUUAGCGGGUUAUCGGGCUAUUCCCGCUAAAAUGGCUAUAGCCAGCUAGAAUGGCGUUUUUAUUA ..((((((......((((((((.............))))))))..((((....))))...))))))............ ( -20.52, z-score = -1.43, R) >droEre2.scaffold_4929 5606040 78 + 26641161 CACGUUUUCAUCCAUUAGCGGAUUAUCGAGCCAUUCCCGCUAAAAUAGCUAUAGCCAGCUAUAAUGGUGGGUUCAUUA ...((((..((((......))))....))))....(((((((..((((((......))))))..)))))))....... ( -20.40, z-score = -2.11, R) >droYak2.chr2L 8649445 62 - 22324452 CACAUGUUUAUUACGUAGCGUGAUAUCGAGCCAUUUCCACUAAAAUAGUUAUAGCCAAAGGU---------------- ..((((((........))))))......(((.((((......)))).)))............---------------- ( -5.10, z-score = 1.12, R) >droSec1.super_5 3598413 74 + 5866729 CACAUUUUCACCAAUUAGCGGGUUAUCGAGCUAUUCCCGAUAAAAUGGCAAUAGCCAGCUAGAAUACUGUUAUU---- ..............(((((.((((((...((((((........)))))).)))))).)))))............---- ( -16.10, z-score = -1.83, R) >droSim1.chr2L 5322593 78 + 22036055 CAUAUUUUCACCAAUUAGCGGGUUAUCGAGCUAUUCCCGCUAAAAUGGCUAUAGCCAGCUAGAAUGGUCACAUCAUUC .........((((.((((((((.............))))))))..((((....)))).......)))).......... ( -21.02, z-score = -2.89, R) >consensus CACAUUUUCACCAAUUAGCGGGUUAUCGAGCUAUUCCCGCUAAAAUGGCUAUAGCCAGCUAGAAUGGUGUUAUCAUU_ ...............((((.((((((..(((((((........))))))))))))).))))................. ( -9.48 = -10.84 + 1.36)



| Location | 5,519,909 – 5,519,987 |
|---|---|
| Length | 78 |
| Sequences | 5 |
| Columns | 78 |
| Reading direction | reverse |
| Mean pairwise identity | 72.68 |
| Shannon entropy | 0.49664 |
| G+C content | 0.40642 |
| Mean single sequence MFE | -17.59 |
| Consensus MFE | -10.10 |
| Energy contribution | -10.58 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.845050 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
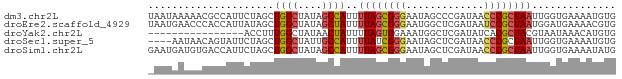
>dm3.chr2L 5519909 78 - 23011544 UAAUAAAAACGCCAUUCUAGCUGGCUAUAGCCAUUUUAGCGGGAAUAGCCCGAUAACCCGCUAAUUGGUGAAAAUGUG .........(((((.......((((....))))..((((((((.............)))))))).)))))........ ( -23.02, z-score = -2.63, R) >droEre2.scaffold_4929 5606040 78 - 26641161 UAAUGAACCCACCAUUAUAGCUGGCUAUAGCUAUUUUAGCGGGAAUGGCUCGAUAAUCCGCUAAUGGAUGAAAACGUG .........((((((..((((.(..((((((((((((....)))))))))..)))..).)))))))).))........ ( -17.80, z-score = -0.98, R) >droYak2.chr2L 8649445 62 + 22324452 ----------------ACCUUUGGCUAUAACUAUUUUAGUGGAAAUGGCUCGAUAUCACGCUACGUAAUAAACAUGUG ----------------....((((((((..((((....))))..))))).)))(((.(((...))).)))........ ( -7.70, z-score = 0.37, R) >droSec1.super_5 3598413 74 - 5866729 ----AAUAACAGUAUUCUAGCUGGCUAUUGCCAUUUUAUCGGGAAUAGCUCGAUAACCCGCUAAUUGGUGAAAAUGUG ----.....((.((...((((((((....))))..((((((((.....))))))))...))))..)).))........ ( -18.80, z-score = -1.97, R) >droSim1.chr2L 5322593 78 - 22036055 GAAUGAUGUGACCAUUCUAGCUGGCUAUAGCCAUUUUAGCGGGAAUAGCUCGAUAACCCGCUAAUUGGUGAAAAUAUG ........(.((((.......((((....))))..((((((((.............)))))))).)))).)....... ( -20.62, z-score = -1.93, R) >consensus _AAUAAAAACACCAUUCUAGCUGGCUAUAGCCAUUUUAGCGGGAAUAGCUCGAUAACCCGCUAAUUGGUGAAAAUGUG .....................((((....))))..((((((((.............)))))))).............. (-10.10 = -10.58 + 0.48)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:19:17 2011