
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 17,304,630 – 17,304,721 |
| Length | 91 |
| Max. P | 0.630064 |

| Location | 17,304,630 – 17,304,721 |
|---|---|
| Length | 91 |
| Sequences | 9 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 68.64 |
| Shannon entropy | 0.62493 |
| G+C content | 0.51509 |
| Mean single sequence MFE | -30.92 |
| Consensus MFE | -9.49 |
| Energy contribution | -9.19 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.54 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.630064 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
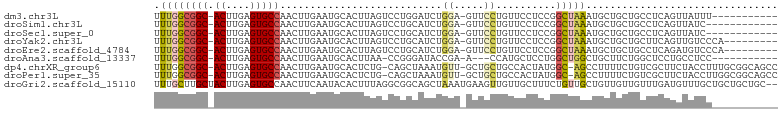
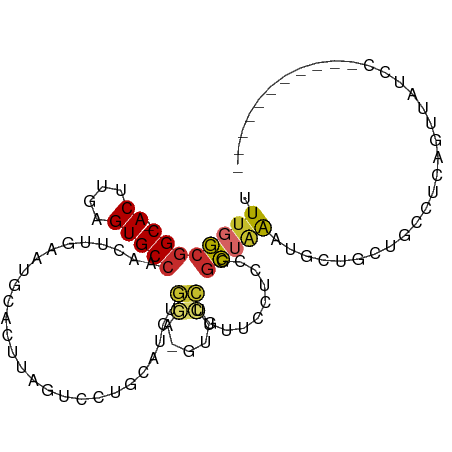
>dm3.chr3L 17304630 91 - 24543557 UUUGGCGGC-ACUUGAGUGCCAACUUGAAUGCACUUAGUCCUGGAUCUGGA-GUUCCUGUUCCUCCGGCUAAAUGCUGCUGCCUCAGUUAUUU----------- ...((((((-(.(..(((....)))..)..(((.((((((..(((...((.-...))...)))...)))))).))))))))))..........----------- ( -31.70, z-score = -2.73, R) >droSim1.chr3L 16641467 90 - 22553184 UUUGGCGGC-ACUUGAGUGCCAACUUGAAUGCACUUAGUCCUGCAUCUGGA-GUUCCUGUUCCUCCGGCUAAAUGCUGCUGCCUCAGUUAUC------------ ...((((((-(.((((((((..........))))))))....(((((((((-(.........))))))....))))))))))).........------------ ( -29.20, z-score = -2.17, R) >droSec1.super_0 9355576 90 - 21120651 UUUGGCGGC-ACUUGAGUGCCAACUUGAAUGCACUUAGUCCUGCAUCUGGA-GUUCCUGUUCCUCCGGCUAAAUGCUGCUGCCUCAGUUAUC------------ ...((((((-(.((((((((..........))))))))....(((((((((-(.........))))))....))))))))))).........------------ ( -29.20, z-score = -2.17, R) >droYak2.chr3L 7431076 93 + 24197627 UUUGGCGGC-ACUUGAGUGCCAACUUGAAUGCACUUAGUCCUGCAUCUGGA-GUUCCUGUUCCUCCGGCUAAAUGCUGCUGCUUCAGUUGUCCCA--------- ...((((((-(.((((((((..........))))))))....(((((((((-(.........))))))....)))))))))))............--------- ( -26.60, z-score = -0.75, R) >droEre2.scaffold_4784 9216899 93 + 25762168 UUUGGCGGC-ACUUGAGUGCCAACUUGAAUGCACUUAGUCCUGCAUCUGGA-GUUCCUGUUCCUCCGGCUAAAUGCUGCUGCCUCAGAUGUCCCA--------- ...((((((-(.((((((((..........))))))))....(((((((((-(.........))))))....)))))))))))............--------- ( -29.20, z-score = -1.58, R) >droAna3.scaffold_13337 8261728 87 - 23293914 UUUGGCGGC-ACUUGAGUGCCAACUUGAAUGCACUUAA-CCGGGAUACCGA-A---CCAUGCUCCUGGCUGGCUGCUUCUGGCUCCUGCCUCC----------- ...((((((-..((((((((..........))))))))-((((((......-.---......))))))...))))))...(((....)))...----------- ( -27.92, z-score = -1.65, R) >dp4.chrXR_group6 3972820 100 + 13314419 UUUGGCGGC-ACUUGAGUGCCAACUUGAAUGCACUCUG-CAGCUAAAUGUU-GCUGCUGCCACUAUGGC-AGCCUUUUCUGUCGCUUCUACCUUUGCGGCAGCC ...((((((-....((((((..........)))))).(-((((........-)))))((((.....)))-)))).....((((((..........))))))))) ( -38.60, z-score = -2.74, R) >droPer1.super_35 384104 100 - 973955 UUUGGCGGC-ACUUGAGUGCCAACUUGAAUGCACUCUG-CAGCUAAAUGUU-GCUGCUGCCACUAUGGC-AGCCUUUUCUGUCGCUUCUACCUUGGCGGCAGCC ...((((((-....((((((..........)))))).(-((((........-)))))((((.....)))-)))).....(((((((........)))))))))) ( -40.40, z-score = -2.81, R) >droGri2.scaffold_15110 13118632 102 - 24565398 UUUGCUUGCUACUUGAGUGCCAACUUCAAUACACUUUAGGCGGCAGCUAAAUGAAGUUGUUGCUUUCUGUUGCUGUUGUUGUUUGAUGUUUGCUGCUGCUGC-- ...((..((.(((..(..(.((((..(((((......(((((((((((......)))))))))))..)))))..)))).)..)..).))..)).))......-- ( -25.50, z-score = 0.14, R) >consensus UUUGGCGGC_ACUUGAGUGCCAACUUGAAUGCACUUAGUCCUGCAUCUGGA_GUUCCUGUUCCUCCGGCUAAAUGCUGCUGCCUCAGUUAUCC___________ ...(((((......((((((..........))))))............((.....)).....................)))))..................... ( -9.49 = -9.19 + -0.30)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:33:23 2011