
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 17,234,643 – 17,234,742 |
| Length | 99 |
| Max. P | 0.957809 |

| Location | 17,234,643 – 17,234,742 |
|---|---|
| Length | 99 |
| Sequences | 14 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 68.21 |
| Shannon entropy | 0.63891 |
| G+C content | 0.51982 |
| Mean single sequence MFE | -33.66 |
| Consensus MFE | -12.24 |
| Energy contribution | -11.95 |
| Covariance contribution | -0.29 |
| Combinations/Pair | 1.55 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.957809 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 17234643 99 - 24543557 CGUGGUCAAACAGAUCCAGCUGCUGUGCAAAAAGCCGCCGAAGGGAUUUGAAAAGUAUUUCGA----AGCCGGCGGCAAGUCGUCUGGCCAACCGAAGGGAUC--------------- ..((((((.((.(((...(((((((.((.........((....)).(((((((....))))))----))))))))))..))))).)))))).((....))...--------------- ( -35.50, z-score = -2.00, R) >droSim1.chr3L 16569544 99 - 22553184 CGUGGUCAAACAGAUCCAGCUGCUGUGCAAGAAGCCGCCGAAGGGAUUCGAAAAGUAUUUCGA----AGCCGGCGGCAAGUCGUCUGGCCAACCGAAGGGAUC--------------- ..((((((.((.(((...(((((((.((.........((....)).(((((((....))))))----))))))))))..))))).)))))).((....))...--------------- ( -37.60, z-score = -2.38, R) >droSec1.super_0 9286698 99 - 21120651 CGUGGUCAAACAGAUCCAGCUGCUGUGCAAGAAGCCGCCGAAGGGAUUCGAAAAGUAUUUCGA----AGCCGGCGGCAAGUCGUCUGGCCAACCGAAGGGAUC--------------- ..((((((.((.(((...(((((((.((.........((....)).(((((((....))))))----))))))))))..))))).)))))).((....))...--------------- ( -37.60, z-score = -2.38, R) >droYak2.chr3L 7359701 99 + 24197627 UGUGGUCAAACAGAUCCAGUUGCUCUGCAAAAAGCCGCCAAAAGGAUUCGAGAAAUAUUUCGA----AGCCGGCGGCAAGUCGUCUGGUCAACCGAAAGGAUC--------------- ..((..((.((.(((....((((...))))...((((((....((.(((((((....))))))----).))))))))..))))).))..)).((....))...--------------- ( -33.70, z-score = -3.05, R) >droEre2.scaffold_4784 9147670 99 + 25762168 UGUGGUCAAACAGAUCCAGCUGCUCUGCAAAAAGCCGCCAAAAGGAUUCGAAAAAUAUUUCGA----AGCCGGCGGCAAGUCGUCUGGCCAACCGAAAGGAUC--------------- ..((((((.((.(((...((......)).....((((((....((.(((((((....))))))----).))))))))..))))).)))))).((....))...--------------- ( -37.50, z-score = -4.01, R) >droAna3.scaffold_13337 7357616 96 - 23293914 CAUUGCCAAACAGAUCCAGCUUCUCUGCAAGAAGCCUCCCAAGGGAUUCGAGAAAUACUUCGA----AGCCGGGGGCAAGUCGUC---ACAACCAAAAGGUGC--------------- ..(((((...........((((((.....))))))..(((...((.((((((......)))))----).))))))))))......---...(((....)))..--------------- ( -26.60, z-score = -0.86, R) >dp4.chrXR_group6 403966 96 + 13314419 CUUCGUGGAGCAGCUGCAGUUGUUCUGCAAGAAGCCGCCGAAGGGCUUUGAAAAGUUCUUUGA----GACCGGCGGCAAGUCCAC---ACAGCCGAAAGGUGC--------------- ....(((((((((..((....)).)))).....(((((((((((((((....))))))))...----...)))))))...)))))---...(((....)))..--------------- ( -40.30, z-score = -2.56, R) >droPer1.super_33 33277 96 + 967471 CUUCGUGGAGCAGCUGCAGUUGUUCUGCAAGAAGCCGCCGAAGGGCUUUGAAAAGUUCUUUGA----GACCGGCGGCAAGUCCAC---ACAGCCGAAAGGUGC--------------- ....(((((((((..((....)).)))).....(((((((((((((((....))))))))...----...)))))))...)))))---...(((....)))..--------------- ( -40.30, z-score = -2.56, R) >droWil1.scaffold_181009 447852 90 + 3585778 CAUAGUCCAACAGAUGCAAUUGUUUUGCAAAAAACCGCCAAAAGGUUUUGAGAAAUAUUUCGA----GGCUGCCGGCAAAAC------ACAACCAAAGGG------------------ ......((......(((((.....))))).......(((....((((((((((....))))))----))))...))).....------.........)).------------------ ( -20.40, z-score = -1.04, R) >droVir3.scaffold_13049 20378307 96 + 25233164 CAUUGUGGAGCAGCUGCAAUUGUUCUGCAAGAAGCCGCCCAAGGGCUUUGAAAAGUAUUUUGA----GGCUGGCGGCAAAUCUGG---CAAACCCAAAGCCGC--------------- ..(((..((((((......))))))..)))...((((((....(((((..(((....)))..)----))))))))))......((---(.........)))..--------------- ( -35.60, z-score = -1.35, R) >droMoj3.scaffold_6680 21517873 93 + 24764193 CAUUGUGGAACAGCUUCAAAUGUUCUGCAAGAAACCGCCGAAGGGUUUUGAAAAGUAUUUCGA----GGCCGGUGGCAAA---GC---UAAACCAAAAGCAGC--------------- ..(((..(((((........)))))..)))......((((...((((((((((....))))))----))))..))))...---((---(........)))...--------------- ( -30.10, z-score = -2.61, R) >droGri2.scaffold_15110 10074584 93 - 24565398 CCUUGUGGAACAGUUUCAAAUGUUUUGCAAGAAGCCGCCCAAGGGUUUUGAGAAGUAUUUCGA----GGCCGGUGGCAAGUCCAC---AAAACCCAAAGC------------------ .((((..(((((........)))))..))))..((((((....((((((((((....))))))----))))))))))........---............------------------ ( -33.20, z-score = -3.41, R) >anoGam1.chr2L 39641970 89 - 48795086 -------------AUACAGGUGUUCUGCGAGAAGCCACCGAAAGGGUUCGAGAAGUACUUCAAA-CCACCCGGCGCCGCUAGCGCACAGCAGGCGAAGCAGGC--------------- -------------..........(((((.....(((..(....)((((.(((......))).))-))....)))..((((.((.....)).))))..))))).--------------- ( -27.10, z-score = -0.18, R) >triCas2.ChLG3 28431152 117 - 32080666 CGUUCUAAACCAGUGGAGACUCUUUUGCGAAAAGCCUCCUAAAGGUUUCGAAAAAUAUUUCGAACCCGGCUCGAAACAGAAGGGCA-AAAGUUCGAAAACUGAGCCUCCGAAGGAGGC ..(((((((..((.(....).)))))).)))..(((((((...((.(((((((....))))))).))((((((........((((.-...))))......)))))).....))))))) ( -35.74, z-score = -1.76, R) >consensus CGUGGUCAAACAGAUCCAGCUGUUCUGCAAGAAGCCGCCGAAGGGAUUCGAAAAGUAUUUCGA____AGCCGGCGGCAAGUCGUC___ACAACCGAAAGGAGC_______________ .................................((((((........(((((......)))))........))))))......................................... (-12.24 = -11.95 + -0.29)


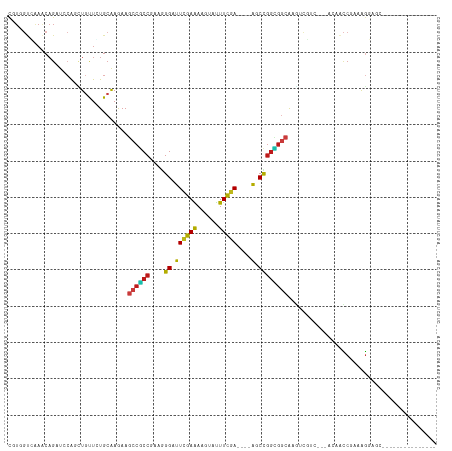
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:33:13 2011