
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 17,176,162 – 17,176,256 |
| Length | 94 |
| Max. P | 0.899110 |

| Location | 17,176,162 – 17,176,256 |
|---|---|
| Length | 94 |
| Sequences | 8 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 71.16 |
| Shannon entropy | 0.51989 |
| G+C content | 0.49684 |
| Mean single sequence MFE | -18.79 |
| Consensus MFE | -10.55 |
| Energy contribution | -10.80 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.899110 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
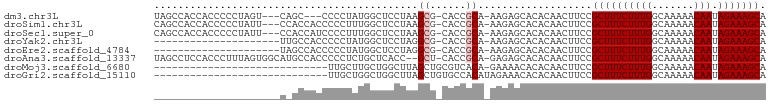
>dm3.chr3L 17176162 94 - 24543557 UAGCCACCACCCCCUAGU---CAGC---CCCCUAUGGCUCCUAAGCG-CACCGCA-AAGAGCACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA ..(((.........(((.---.(((---(......)))).))).(((-...))).-..).))...........((((((((((.......))).))))))). ( -19.80, z-score = -1.66, R) >droSim1.chr3L 16510528 97 - 22553184 CAGCCACCACCCCCUAUU---CCACCACCCCCUUUGGCUCCUAAGCG-CACCGCA-AAGAGCACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA ..................---.............((((((....(((-...))).-..)))).))........((((((((((.......))).))))))). ( -17.50, z-score = -1.93, R) >droSec1.super_0 9228132 97 - 21120651 CAGCCACCACCCCCUAUU---CCACCAUCCCCUUUGGCUCCUAAGCG-CACCGCA-AAGAGCACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA ..................---.............((((((....(((-...))).-..)))).))........((((((((((.......))).))))))). ( -17.50, z-score = -1.86, R) >droYak2.chr3L 7298174 79 + 24197627 ---------------------UUGCCACCCCCUAUGGCUCCUAGGCG-CACCGCA-AAGAGCACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA ---------------------.............((((((....(((-...))).-..)))).))........((((((((((.......))).))))))). ( -18.70, z-score = -1.23, R) >droEre2.scaffold_4784 9087283 79 + 25762168 ---------------------UAGCCACCCCCUAUGGCUCCUAGGCG-CACCGCA-AAGAGCACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA ---------------------.............((((((....(((-...))).-..)))).))........((((((((((.......))).))))))). ( -18.70, z-score = -1.37, R) >droAna3.scaffold_13337 7299697 98 - 23293914 UAGCCUCCACCCUUUAGUGGCAUGCCACCCCCUCUGCUCACC--GCU-CACCGCA-GAGAGCACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA ..((............((((....))))...((((((.....--...-....)))-))).))...........((((((((((.......))).))))))). ( -25.70, z-score = -3.00, R) >droMoj3.scaffold_6680 21448782 72 + 24764193 -----------------------------UUGCUUGCUGGCUUAGCUGCGUCACA-GAAAACACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA -----------------------------..((..((((...)))).))......-.................((((((((((.......))).))))))). ( -14.60, z-score = 0.01, R) >droGri2.scaffold_15110 10013355 73 - 24565398 -----------------------------UUGCUGGCUGGCUUAGCUGUGCCACAUAGAAACACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA -----------------------------.((.((((((((...)))).))))))..................((((((((((.......))).))))))). ( -17.80, z-score = -0.79, R) >consensus _____________________C_GCCACCCCCUUUGGCUCCUAAGCG_CACCGCA_AAGAGCACACAACUUCCGCUUUCUUUGGCAAAAACAAUAGAAAGCA ............................................((......))...................((((((((((.......))).))))))). (-10.55 = -10.80 + 0.25)

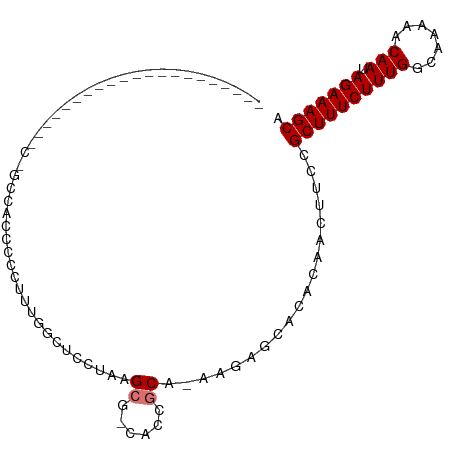
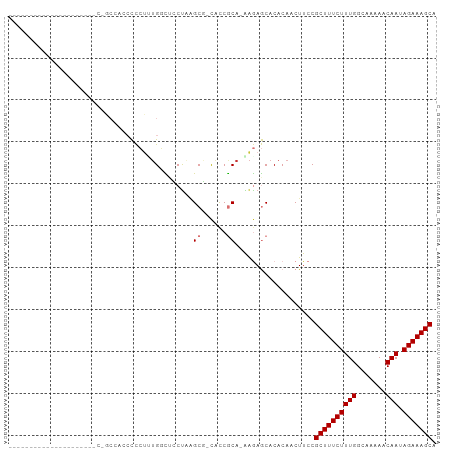
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:33:05 2011