
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 17,110,762 – 17,110,855 |
| Length | 93 |
| Max. P | 0.830024 |

| Location | 17,110,762 – 17,110,855 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 72.40 |
| Shannon entropy | 0.55943 |
| G+C content | 0.48131 |
| Mean single sequence MFE | -16.43 |
| Consensus MFE | -10.75 |
| Energy contribution | -10.99 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.06 |
| Mean z-score | -0.82 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.830024 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 17110762 93 - 24543557 -----CUCAGCCCAACUCAAACCCCUCGA-------------CCAACCCAUCGUGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAUC -----...(((............((.(((-------------........))).))......(((.(((((((((........)))).))))).)))....)))....... ( -13.20, z-score = -1.19, R) >droSim1.chr3L 16445922 108 - 22553184 CAGACCUCAGCC---CUCAAACCCCUCGAACAACCCAUCGCCCCAACCCAGCGUGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGUUCAUUUUCGUCGCUUAUCAUC ............---............(((((...((.(((.........)))))((((((....))))))................)))))................... ( -13.10, z-score = 0.00, R) >droSec1.super_0 9161554 108 - 21120651 CAGACCUCAGCC---CUCAAACCCCUCGAACAACGCAUCGCCCCAACCCAGCGUGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAUC ..(((....(((---.((.........))...((((..............))))))).....(((.(((((((((........)))).))))).))).))).......... ( -16.54, z-score = -0.43, R) >droYak2.chr3L 7229486 92 + 24197627 ----------------UCGAUUCCCCCGAACCACCCUUC---CCAGCCCAUCGUGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAUC ----------------..(((..(..((((.........---...(((......))).........(((((((((........)))).)))))..))))..)...)))... ( -15.70, z-score = -1.83, R) >droEre2.scaffold_4784 9020852 92 + 25762168 ----------------CCAAUGCUCUCGAACUACCCAUC---CCCACCCAUCGUGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAUC ----------------.....((...((((.........---.((((.....))))..........(((((((((........)))).)))))..))))..))........ ( -15.40, z-score = -2.12, R) >droAna3.scaffold_13337 7234246 89 - 23293914 ----------------UCGCUGCUUGCACGCUGCCC-----CCCAACCCCAU-UGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAUC ----------------..((.....))..((((((.-----...........-.))))....(((.(((((((((........)))).))))).)))....))........ ( -14.32, z-score = -0.10, R) >dp4.chrXR_group6 13237947 104 - 13314419 ---CCGCCCGCCGCCCAUUCAACACUCGGCUUUCGCUU---UGCCUUCUG-UGUGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAUC ---..((..(((((.......(((...(((........---.)))...))-)))))).....(((.(((((((((........)))).))))).)))....))........ ( -20.51, z-score = -0.51, R) >droPer1.super_250 902 102 - 39459 -----GCCCGCCCCCCAUUCAACACUCGGCUUUCGCUU---UGCCUUCUG-UGUGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAUC -----((..(((.........((((..(((........---.)))....)-)))))).....(((.(((((((((........)))).))))).)))....))........ ( -18.70, z-score = -0.74, R) >droWil1.scaffold_181009 3294258 100 - 3585778 --------CACCACGCCACCGUUUCUGCUACUUCCGGU---UGCCUCCUACUGGGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAGC --------......((.((.(((((.......((((((---........))))))((((((....))))))))))).....((.....))........)).))........ ( -20.40, z-score = -0.44, R) >consensus _______C_GCC___CUCAAACCCCUCGAACUACCCAU____CCAACCCAUCGUGGCAUAAAAAAUUGUGCGAAACUUUCAGCUUUCUGCACAUUUUCGUCGCUUAUCAUC ....................................................(((((....(((((.((((((((........)))).))))))))).)))))........ (-10.75 = -10.99 + 0.24)


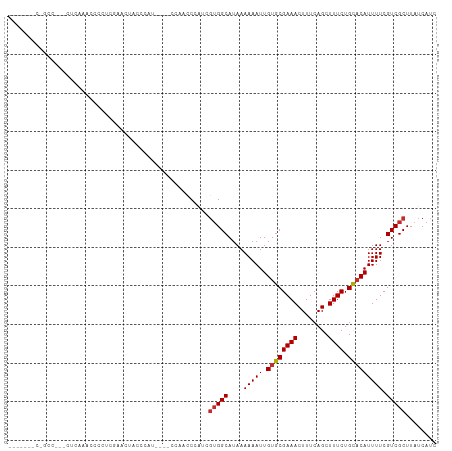
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:32:53 2011