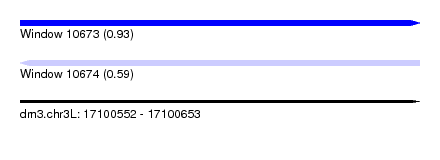

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 17,100,552 – 17,100,653 |
| Length | 101 |
| Max. P | 0.931820 |

| Location | 17,100,552 – 17,100,653 |
|---|---|
| Length | 101 |
| Sequences | 8 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 79.34 |
| Shannon entropy | 0.41000 |
| G+C content | 0.44807 |
| Mean single sequence MFE | -31.16 |
| Consensus MFE | -16.71 |
| Energy contribution | -18.01 |
| Covariance contribution | 1.30 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.931820 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
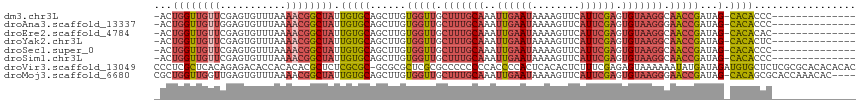
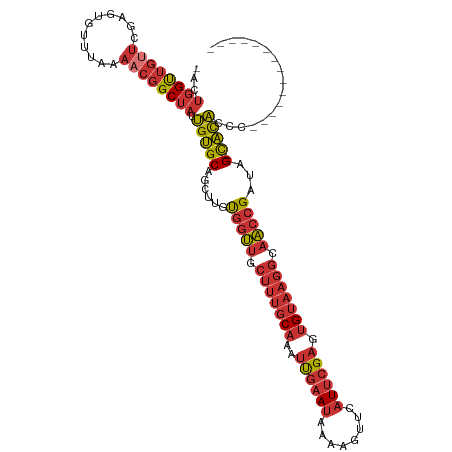
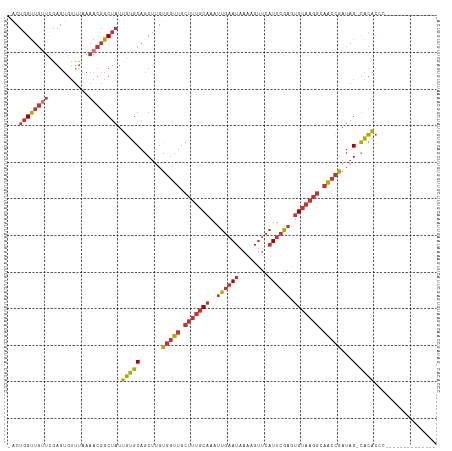
>dm3.chr3L 17100552 101 + 24543557 -ACUGGUUGUUCGAGUGUUUAAAACGGCUAUUGUGCAGCUUGUGGUUGCUUUGCAAAUUGAAUAAAAGUUCAUUCGAGUGUAAGGCAACCGAUAG-CACACCC-------------- -..((((((((...........)))))))).(((((......(((((((((((((..((((((........)))))).)))))))))))))...)-))))...-------------- ( -32.00, z-score = -2.51, R) >droAna3.scaffold_13337 7224409 101 + 23293914 -ACUGGUUGUUGGAGUGUUUAAAACGGCUAUUGUGCAGCUUGUGGUUGCUUUGCAAAUUGAAUAAAAGUUCAUUCGAGUGUAAGGGAACCGAUAG-CACACCC-------------- -..((((((((...........)))))))).(((((......(((((.(((((((..((((((........)))))).))))))).)))))...)-))))...-------------- ( -28.70, z-score = -1.49, R) >droEre2.scaffold_4784 9010529 101 - 25762168 -ACUGGUUGUUCGAGUGUUUAAAACGGCUAUUGUGCAGCUUGUGGUUGCUUUGCAAAUUGAAUAAAAGUUCAUUCGAGUGUAAGGCAACCGAUAG-CACACAC-------------- -..((((((((...........)))))))).((((..(((..(((((((((((((..((((((........)))))).)))))))))))))..))-).)))).-------------- ( -33.00, z-score = -2.73, R) >droYak2.chr3L 7218697 101 - 24197627 -ACUGGUUGUUCGAGUGUUUAAAACGGCUAUUGUGCAGCUUGUGGUUGCUUUGCAAAUUGAAUAAAAGUUCAUUCGAGUGUAAGGCAACCGAUAG-CACACUC-------------- -...........((((((........((......)).(((..(((((((((((((..((((((........)))))).)))))))))))))..))-)))))))-------------- ( -34.30, z-score = -3.34, R) >droSec1.super_0 9151330 101 + 21120651 -ACUGGUUGUUCGAGUGUUUAAAACGGCUAUUGUGCAGCUUGUGGUUGCUUUGCAAAUUGAAUAAAAGUUCAUUCGAGUGUAAGGCAACCGAUAG-CACACCC-------------- -..((((((((...........)))))))).(((((......(((((((((((((..((((((........)))))).)))))))))))))...)-))))...-------------- ( -32.00, z-score = -2.51, R) >droSim1.chr3L 16433914 101 + 22553184 -ACUGGUUGUUCGAGUGUUUAAAACGGCUAUUGUGCAGCUUGUGGUUGCUUUGCAAAUUGAAUAAAAGUUCAUUCGAGUGUAAGGCAACCGAUAG-CACACCC-------------- -..((((((((...........)))))))).(((((......(((((((((((((..((((((........)))))).)))))))))))))...)-))))...-------------- ( -32.00, z-score = -2.51, R) >droVir3.scaffold_13049 20218148 116 - 25233164 CCCUCGCUCACAGAGACACCACACACGCUCUCGCGC-GCGCGCUCGCGCCCCCCCCACCCCACUCACACUCUUUCGAGAGUAAAAAAUAUGAUAGAUGUGCUCUCGCGCACACACAC ............((((............))))((((-((((....))))..........................(((((((....((.......)).)))))))))))........ ( -25.40, z-score = -2.02, R) >droMoj3.scaffold_6680 21355665 112 - 24764193 CGCUGGUUGGUUGAGUGUUUAAAACGGCUAUUGUGCAGCUUGUGGUUGCUUUGCAAAUUGAAUAAAAGUUCAUUCGAGUGUAAGGGAACCGAUAG-CACAGCGCACCAAACAC---- ...((((((((((..(.....)..))))))..((((.(((..(((((.(((((((..((((((........)))))).))))))).)))))..))-)...)))))))).....---- ( -31.90, z-score = -0.77, R) >consensus _ACUGGUUGUUCGAGUGUUUAAAACGGCUAUUGUGCAGCUUGUGGUUGCUUUGCAAAUUGAAUAAAAGUUCAUUCGAGUGUAAGGCAACCGAUAG_CACACCC______________ ...((((((((...........)))))))).((((.......(((((.(((((((..((((((........)))))).))))))).))))).....))))................. (-16.71 = -18.01 + 1.30)
| Location | 17,100,552 – 17,100,653 |
|---|---|
| Length | 101 |
| Sequences | 8 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 79.34 |
| Shannon entropy | 0.41000 |
| G+C content | 0.44807 |
| Mean single sequence MFE | -25.96 |
| Consensus MFE | -14.44 |
| Energy contribution | -16.62 |
| Covariance contribution | 2.19 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.589417 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
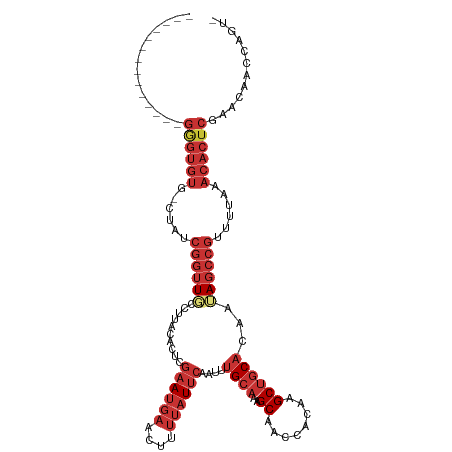
>dm3.chr3L 17100552 101 - 24543557 --------------GGGUGUG-CUAUCGGUUGCCUUACACUCGAAUGAACUUUUAUUCAAUUUGCAAAGCAACCACAAGCUGCACAAUAGCCGUUUUAAACACUCGAACAACCAGU- --------------((((((.-.....((((((.((.((...((((((....))))))....)).)).))))))..((((.((......)).))))...))))))...........- ( -23.40, z-score = -1.51, R) >droAna3.scaffold_13337 7224409 101 - 23293914 --------------GGGUGUG-CUAUCGGUUCCCUUACACUCGAAUGAACUUUUAUUCAAUUUGCAAAGCAACCACAAGCUGCACAAUAGCCGUUUUAAACACUCCAACAACCAGU- --------------((((((.-....(((((...........((((((....))))))....((((..((........))))))....)))))......)))).))..........- ( -20.20, z-score = -1.24, R) >droEre2.scaffold_4784 9010529 101 + 25762168 --------------GUGUGUG-CUAUCGGUUGCCUUACACUCGAAUGAACUUUUAUUCAAUUUGCAAAGCAACCACAAGCUGCACAAUAGCCGUUUUAAACACUCGAACAACCAGU- --------------.((((((-((...((((((.((.((...((((((....))))))....)).)).))))))...))).)))))......((((.........)))).......- ( -20.60, z-score = -0.79, R) >droYak2.chr3L 7218697 101 + 24197627 --------------GAGUGUG-CUAUCGGUUGCCUUACACUCGAAUGAACUUUUAUUCAAUUUGCAAAGCAACCACAAGCUGCACAAUAGCCGUUUUAAACACUCGAACAACCAGU- --------------((((((.-.....((((((.((.((...((((((....))))))....)).)).))))))..((((.((......)).))))...))))))...........- ( -24.30, z-score = -2.35, R) >droSec1.super_0 9151330 101 - 21120651 --------------GGGUGUG-CUAUCGGUUGCCUUACACUCGAAUGAACUUUUAUUCAAUUUGCAAAGCAACCACAAGCUGCACAAUAGCCGUUUUAAACACUCGAACAACCAGU- --------------((((((.-.....((((((.((.((...((((((....))))))....)).)).))))))..((((.((......)).))))...))))))...........- ( -23.40, z-score = -1.51, R) >droSim1.chr3L 16433914 101 - 22553184 --------------GGGUGUG-CUAUCGGUUGCCUUACACUCGAAUGAACUUUUAUUCAAUUUGCAAAGCAACCACAAGCUGCACAAUAGCCGUUUUAAACACUCGAACAACCAGU- --------------((((((.-.....((((((.((.((...((((((....))))))....)).)).))))))..((((.((......)).))))...))))))...........- ( -23.40, z-score = -1.51, R) >droVir3.scaffold_13049 20218148 116 + 25233164 GUGUGUGUGCGCGAGAGCACAUCUAUCAUAUUUUUUACUCUCGAAAGAGUGUGAGUGGGGUGGGGGGGGCGCGAGCGCGC-GCGCGAGAGCGUGUGUGGUGUCUCUGUGAGCGAGGG ((((....))))..(..(((.((((((......(((((.(((....))).)))))...))))))((((((((...(((((-((((....))))))))))))))))))))..)..... ( -49.70, z-score = -3.31, R) >droMoj3.scaffold_6680 21355665 112 + 24764193 ----GUGUUUGGUGCGCUGUG-CUAUCGGUUCCCUUACACUCGAAUGAACUUUUAUUCAAUUUGCAAAGCAACCACAAGCUGCACAAUAGCCGUUUUAAACACUCAACCAACCAGCG ----(((((((((..(((.((-(.....((......))....((((((....)))))).....))).))).)))..((((.((......)).)))).)))))).............. ( -22.70, z-score = -0.34, R) >consensus ______________GGGUGUG_CUAUCGGUUGCCUUACACUCGAAUGAACUUUUAUUCAAUUUGCAAAGCAACCACAAGCUGCACAAUAGCCGUUUUAAACACUCGAACAACCAGU_ ..............((((((......((((((..........((((((....))))))....((((..((........))))))...))))))......))))))............ (-14.44 = -16.62 + 2.19)

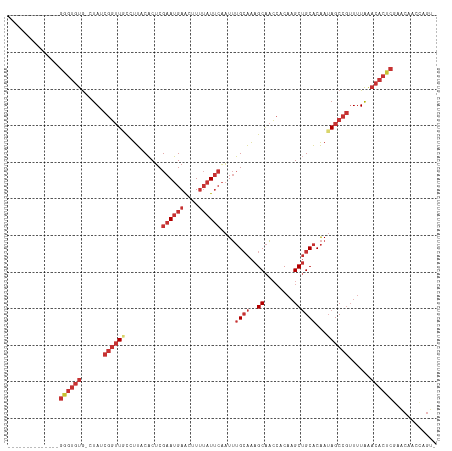
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:32:52 2011