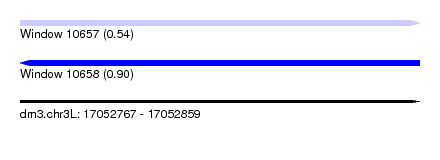

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 17,052,767 – 17,052,859 |
| Length | 92 |
| Max. P | 0.904332 |

| Location | 17,052,767 – 17,052,859 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 90.88 |
| Shannon entropy | 0.14002 |
| G+C content | 0.32962 |
| Mean single sequence MFE | -20.70 |
| Consensus MFE | -15.45 |
| Energy contribution | -15.45 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.535373 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
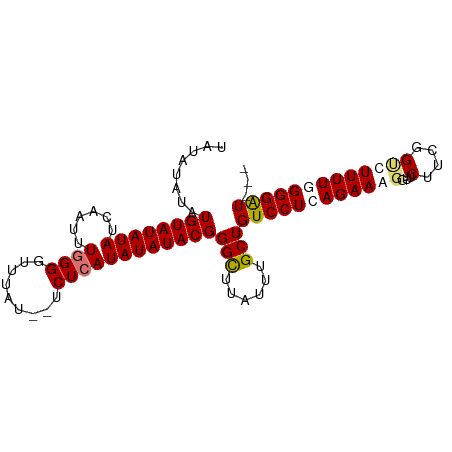
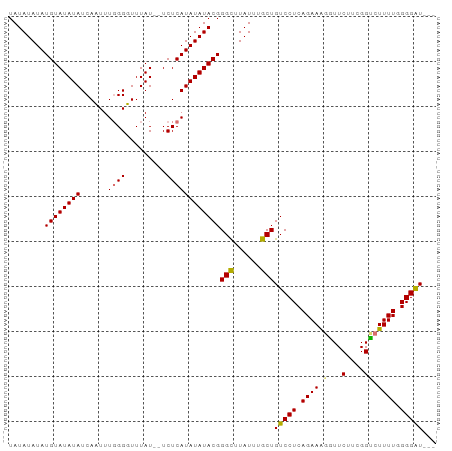
>dm3.chr3L 17052767 92 + 24543557 UAUAUAUAUGUAUAUAUCAAUUUGGGUUUUAUUUUCUCAUAUAUACGGGCUUAUUUGCUGUCCUCAGAAGGGUUCUUCGGCCUUUUUGGGAU--- ........((((((((......((((.........))))))))))))(((......)))((((.(((((((((......)))))))))))))--- ( -23.30, z-score = -2.57, R) >droSec1.super_0 9113164 92 + 21120651 UAUAUAUAUGUAUAUAUCAAUUUGGGUUUUAUUUUCUCAUAUAUACGGGCUUAUUUGCUGUCCUCAGAAGGAUUCUUCGGUCUUUUUGGGAU--- ........((((((((......((((.........))))))))))))(((......)))((((.(((((((((......)))))))))))))--- ( -21.30, z-score = -2.39, R) >droYak2.chr3L 7178477 93 - 24197627 UAUAUAUAUGUAUAUAUCAUUUUGGGGUUUAU--UCUCAUAUAUACGGGCUUAUUUGCUGUCCUCAGAAAUGUUCUUCGGUCUUUUGGGGGUUCU ........((((((((......(((((.....--)))))))))))))(((......))).(((((((((.....(....)..))))))))).... ( -19.40, z-score = -1.49, R) >droEre2.scaffold_4784 8972654 90 - 25762168 UAUAUAUAUGUAUAUAUCAUUUUUGGGUUUAU--UCUCAUAUAUACGGGUUUAUUUGCUGUCCUCAGAAAGGUUCUUCGGUCUUUUGGGGAU--- ..((((((((.(((..(((....)))...)))--...))))))))..(((......)))((((((((((.((..(....)))))))))))))--- ( -18.80, z-score = -1.61, R) >consensus UAUAUAUAUGUAUAUAUCAAUUUGGGGUUUAU__UCUCAUAUAUACGGGCUUAUUUGCUGUCCUCAGAAAGGUUCUUCGGUCUUUUGGGGAU___ ........((((((((......((((.........))))))))))))(((......)))((((.(((((((((......)))))))))))))... (-15.45 = -15.45 + 0.00)

| Location | 17,052,767 – 17,052,859 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 90.88 |
| Shannon entropy | 0.14002 |
| G+C content | 0.32962 |
| Mean single sequence MFE | -10.34 |
| Consensus MFE | -9.20 |
| Energy contribution | -9.21 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.89 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.904332 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
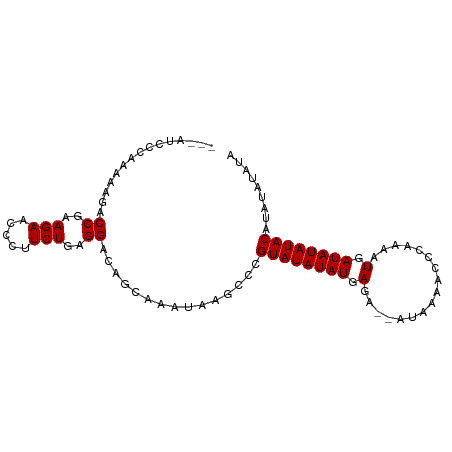
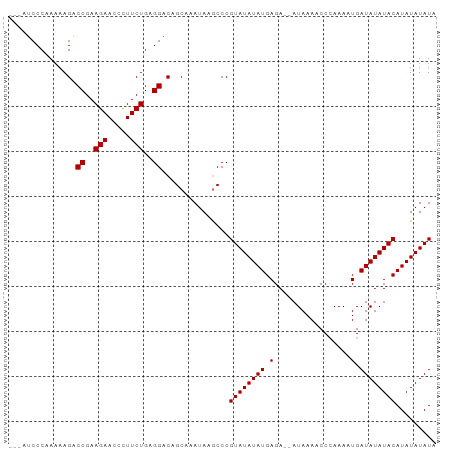
>dm3.chr3L 17052767 92 - 24543557 ---AUCCCAAAAAGGCCGAAGAACCCUUCUGAGGACAGCAAAUAAGCCCGUAUAUAUGAGAAAAUAAAACCCAAAUUGAUAUAUACAUAUAUAUA ---..........((((..(((.....)))..))...((......))))((((((((.((...............)).))))))))......... ( -10.86, z-score = -1.15, R) >droSec1.super_0 9113164 92 - 21120651 ---AUCCCAAAAAGACCGAAGAAUCCUUCUGAGGACAGCAAAUAAGCCCGUAUAUAUGAGAAAAUAAAACCCAAAUUGAUAUAUACAUAUAUAUA ---............((..(((.....)))..))...((......))..((((((((.((...............)).))))))))......... ( -9.86, z-score = -1.46, R) >droYak2.chr3L 7178477 93 + 24197627 AGAACCCCCAAAAGACCGAAGAACAUUUCUGAGGACAGCAAAUAAGCCCGUAUAUAUGAGA--AUAAACCCCAAAAUGAUAUAUACAUAUAUAUA ...............((..(((.....)))..))...((......))..((((((((.(..--.............).))))))))......... ( -10.56, z-score = -2.82, R) >droEre2.scaffold_4784 8972654 90 + 25762168 ---AUCCCCAAAAGACCGAAGAACCUUUCUGAGGACAGCAAAUAAACCCGUAUAUAUGAGA--AUAAACCCAAAAAUGAUAUAUACAUAUAUAUA ---............((..(((.....)))..))...............((((((((.(..--.............).))))))))......... ( -10.06, z-score = -2.27, R) >consensus ___AUCCCAAAAAGACCGAAGAACCCUUCUGAGGACAGCAAAUAAGCCCGUAUAUAUGAGA__AUAAAACCCAAAAUGAUAUAUACAUAUAUAUA ...............((..(((.....)))..))...............((((((((.(.................).))))))))......... ( -9.20 = -9.21 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:32:40 2011