
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 16,635,583 – 16,635,683 |
| Length | 100 |
| Max. P | 0.834455 |

| Location | 16,635,583 – 16,635,683 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 74.36 |
| Shannon entropy | 0.45573 |
| G+C content | 0.37753 |
| Mean single sequence MFE | -24.18 |
| Consensus MFE | -16.33 |
| Energy contribution | -17.75 |
| Covariance contribution | 1.42 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.834455 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
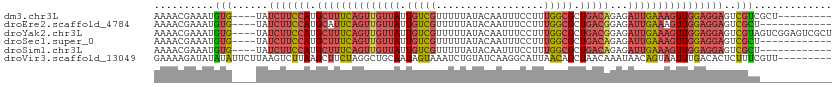
>dm3.chr3L 16635583 100 - 24543557 AAAACGAAAUGUG----UAUCUUCCAUGCUUUCAGUUGUUAUUGUCGUUUUUAUACAAUUUCCUUUGGCGCUGACAGAGAUUGAAAGUUGGAGGAGUCGUCGCU--------- ..........(((----..(((((((.((((((((((((((.(((((..................))))).)))))...)))))))))))))))).....))).--------- ( -26.57, z-score = -1.82, R) >droEre2.scaffold_4784 8569435 97 + 25762168 AAAACGAAAUGUG----UAUCUUCCAUGCAUUCAGUUGUUAUUGUCGUUUUUAUACAAUUUCCUUUGGCGCUGACGGAGAUUGAAAGUUGGAGGAGUCGCU------------ ..........(((----..(((((((.((.(((((((((((.(((((..................))))).))))...))))))).)))))))))..))).------------ ( -23.07, z-score = -0.76, R) >droYak2.chr3L 6762500 109 + 24197627 AAAACGAAAUGUG----UAUCUUCCAUGCUUUCAGUUGUUAUUGUCGUUUUUAUACAAUUUCCUUUGGCGCUGACGGAGAUUGAAAGUUGGAGGAGUCGUAGUCGGAGUCGCU ....(((.(((..----..(((((((.((((((((((((((.(((((..................))))).))))...)))))))))))))))))..)))..)))........ ( -28.17, z-score = -0.96, R) >droSec1.super_0 8713467 97 - 21120651 AAAACGAAAUGUG----UAUCUUCCAUGCUUUCAGUUGUUAUUGUCGUUUUUAUACAAUUUCCUUUGGCGCUGACAGAGAUUGAAAGUUGGAGGAGUCGCU------------ ..........(((----..(((((((.((((((((((((((.(((((..................))))).)))))...))))))))))))))))..))).------------ ( -27.47, z-score = -2.50, R) >droSim1.chr3L 15966337 97 - 22553184 AAAACGAAAUGUG----UAUCUUCCAUGCUUUCAGUUGUUAUUGUCGUUUUUAUACAAUUUCCUUUGGCGCUGACAGAGAUUGAAAGUUGGAGGAGUCGCU------------ ..........(((----..(((((((.((((((((((((((.(((((..................))))).)))))...))))))))))))))))..))).------------ ( -27.47, z-score = -2.50, R) >droVir3.scaffold_13049 3724919 104 - 25233164 GAAAAGAUAUAUAUUCUUAAGUCUUAAGCUUCUAGGCUGCAAUAGUAAAUCUGUAUCAAGGCAUUAACAGCUAACAAAUAACAGUAAUUUGACACUCUUUCGUU--------- ((((((((.((......)).))))).((((..((((((...((((.....)))).....))).)))..))))..(((((.......))))).......)))...--------- ( -12.30, z-score = 1.05, R) >consensus AAAACGAAAUGUG____UAUCUUCCAUGCUUUCAGUUGUUAUUGUCGUUUUUAUACAAUUUCCUUUGGCGCUGACAGAGAUUGAAAGUUGGAGGAGUCGCU____________ ...................(((((((.((((((((((((((.(((((..................))))).)))))...)))))))))))))))).................. (-16.33 = -17.75 + 1.42)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:31:37 2011