
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 16,553,866 – 16,553,964 |
| Length | 98 |
| Max. P | 0.515827 |

| Location | 16,553,866 – 16,553,964 |
|---|---|
| Length | 98 |
| Sequences | 8 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 67.89 |
| Shannon entropy | 0.61172 |
| G+C content | 0.49434 |
| Mean single sequence MFE | -24.50 |
| Consensus MFE | -11.13 |
| Energy contribution | -10.61 |
| Covariance contribution | -0.51 |
| Combinations/Pair | 1.46 |
| Mean z-score | -0.98 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.515827 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
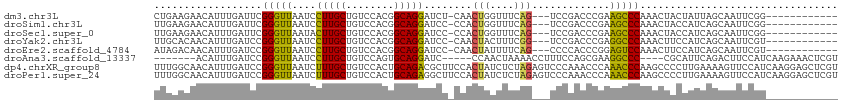
>dm3.chr3L 16553866 98 + 24543557 CUGAAGAACAUUUGAUUCGGGUUAAUCCUUGCUGUCCACGGCAGGAUCU-CAACUGGUUUCAG---UCCGACCCGAAGCCCAAACUACUAUUAGCAAUUCGG------------ (((((.....((((.((((((((.(((((.((((....)))))))))..-..((((....)))---)..))))))))...))))(((....)))...)))))------------ ( -27.50, z-score = -1.91, R) >droSim1.chr3L 15899129 98 + 22553184 UUGAAGAACAUUUGAUUCGGGUUAAUCCUUGCUGUCCACGGCAGGAUCC-CCACUGGUUUCAG---UCCGACCCGAAGCCCAAACUACCAUCAGCAAUUCGG------------ ((((.(....((((.((((((((.(((((.((((....)))))))))..-..((((....)))---)..))))))))...))))....).))))........------------ ( -26.50, z-score = -1.28, R) >droSec1.super_0 8646497 98 + 21120651 UUGAAGAACAUUUGAUUCGGGUUAAUACUUGCUGUCCACGGCAGGAUCC-CCACUGGUUUCAG---UCCGACCCGAAGCCCAAACUACCAUCAGCAAUUCGG------------ ((((.(....((((.((((((((....(((((((....)))))))....-..((((....)))---)..))))))))...))))....).))))........------------ ( -25.80, z-score = -1.23, R) >droYak2.chr3L 6693376 98 - 24197627 UUGCACAACAUUUGAUCCGGGUUAAUCCUUGCUGUCCACGGCAGGAUCC-CAACUACUUUCGG---UCCGACCCGAGGCCCAAACUUCCAUCAGCAAUUCGU------------ ((((......((((..(((((...(((((.((((....)))))))))))-)........((((---......))))))..)))).........)))).....------------ ( -24.36, z-score = -0.90, R) >droEre2.scaffold_4784 8500135 98 - 25762168 AUAGACAACAUUUGAUCCGGGUUAAUCCUUGCUGUCCACGGCAGGAUCC-CAACUAUUUUCAG---CCCCACCCGGAGUCCAAACUUCCAUCAGCAAUUCGU------------ ..........((((.(((((((..(((((.((((....)))))))))..-...((......))---....)))))))...))))..................------------ ( -22.30, z-score = -1.40, R) >droAna3.scaffold_13337 9271630 98 - 23293914 -------ACAUUUGAUCCGGGUUAAUCCUUGCUGUCCAGUGCAGGAUC-----CCAACUAAAACCUUUCCAGCGAAGGCCC----CGCAUUCAGACUUCCAUCAAGAAACUCGU -------...(((((..((((...(((((.((........))))))).-----..........(((((.....))))).))----))...)))))................... ( -19.60, z-score = -0.18, R) >dp4.chrXR_group8 6955373 114 - 9212921 UUUGGCAACAUUUGAUCCGGGUUAAUCUUUGCUGUCCACUGCAGACGCUUCCACUAUCUCUAGAGUCCCAAACCCAAACCCAAGCCCCUUGAAAAGUUCCAUCAAGGAGCUCGU ..((....))........(((((......(((........)))(((.((............)).)))...))))).......(((.((((((.........)))))).)))... ( -23.40, z-score = -0.35, R) >droPer1.super_24 798861 114 + 1556852 UUUGGCAACAUUUGAUCCGGGUUAAUCUUUGCUGUCCACUGCAGAGGCUUCCACUAUCUCUAGAGUCCCAAACCCAAACCCAAGCCCCUUGAAAAGUUCCAUCAAGGAGCUCGU ..((....))........(((((..(((((((........)))))))......(((....))).......))))).......(((.((((((.........)))))).)))... ( -26.50, z-score = -0.63, R) >consensus UUGAACAACAUUUGAUCCGGGUUAAUCCUUGCUGUCCACGGCAGGAUCC_CCACUAGUUUCAG___UCCGACCCGAAGCCCAAACUACCAUCAGCAAUUCGU____________ ..................(((((....(((((........)))))........(((....))).............)))))................................. (-11.13 = -10.61 + -0.51)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:31:25 2011