
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 16,445,708 – 16,445,765 |
| Length | 57 |
| Max. P | 0.599317 |

| Location | 16,445,708 – 16,445,765 |
|---|---|
| Length | 57 |
| Sequences | 10 |
| Columns | 65 |
| Reading direction | reverse |
| Mean pairwise identity | 84.04 |
| Shannon entropy | 0.32097 |
| G+C content | 0.45484 |
| Mean single sequence MFE | -13.04 |
| Consensus MFE | -8.53 |
| Energy contribution | -8.54 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.599317 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
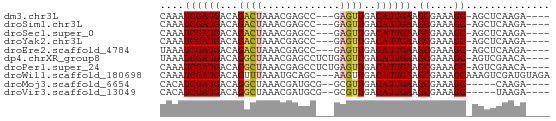
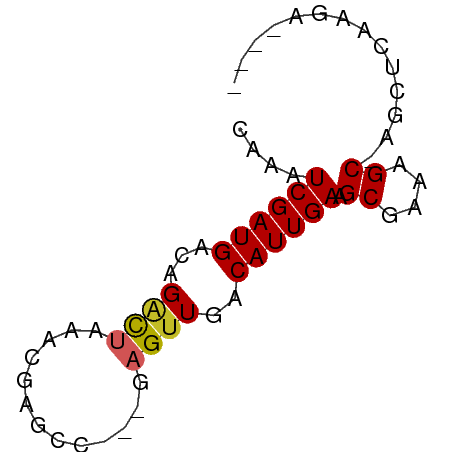
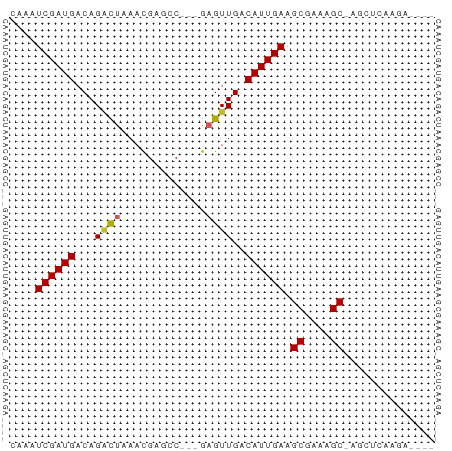
>dm3.chr3L 16445708 57 - 24543557 CAAAUCGAUGACAGACUAAACGAGCC---GAGUUGACAUUGAAGCGAAAGC-AGCUCAAGA---- ....((((((...((((.........---.))))..)))))).((....))-.........---- ( -11.60, z-score = -1.51, R) >droSim1.chr3L 15787307 57 - 22553184 CAAAUCGAUGACAGACUAAACGAGCC---GAGUUGACAUUGAAGCGAAAGC-AGCUCAAGA---- ....((((((...((((.........---.))))..)))))).((....))-.........---- ( -11.60, z-score = -1.51, R) >droSec1.super_0 8537550 57 - 21120651 CAAAUCGAUGACAGACUAAACGAGCC---GAGUUGACAUUGAAGCGAAAGC-AGCUCAAGA---- ....((((((...((((.........---.))))..)))))).((....))-.........---- ( -11.60, z-score = -1.51, R) >droYak2.chr3L 6584494 57 + 24197627 CAAAUCGAUGACAGACUAAACGAGCC---GAGUUGACAUUGAAGCGAAAGC-AGCUCAAGA---- ....((((((...((((.........---.))))..)))))).((....))-.........---- ( -11.60, z-score = -1.51, R) >droEre2.scaffold_4784 8395219 57 + 25762168 UAAAUCGAUGACAGACUAAACGAGCC---GAGUUGACAUUGAAGCGAAAGC-AGCUCAAGA---- ....((((((...((((.........---.))))..)))))).((....))-.........---- ( -11.60, z-score = -1.74, R) >dp4.chrXR_group8 2137564 60 - 9212921 UAAAUCGAUGACAGGCUAAACGAGCCUCUGAGUUGACAUUGAAGCGAAAGC-AGUCGAACA---- ....(((((...(((((.....)))))....)))))..((((.((....))-..))))...---- ( -16.00, z-score = -2.18, R) >droPer1.super_24 636265 60 + 1556852 CAAAUCGAUGACAGGCUAAACGAGCCUCUGAGUUGACAUUGAAGCGAAAGC-AGUCGAACA---- ....(((((...(((((.....)))))....)))))..((((.((....))-..))))...---- ( -16.00, z-score = -1.96, R) >droWil1.scaffold_180698 2989941 62 + 11422946 CAAAUCGAUGACAGUUUAAAUGCAGC---AAGUUGACAUUGAAGCGAAAGCAAAGUCGAUGUAGA ...((((((.........(((((((.---...))).))))...((....))...))))))..... ( -13.80, z-score = -1.79, R) >droMoj3.scaffold_6654 179471 54 + 2564135 CACAUCGAUGACAGGCUAAACGAUGCG--GCGUUGACAUUGAAGCGAAAGC-----CAAGA---- ....((((((.((((((.........)--)).))).)))))).((....))-----.....---- ( -12.50, z-score = -1.16, R) >droVir3.scaffold_13049 1439641 54 - 25233164 CACAUCGAUGACAGGCUAAACGAUGCG--GCGUUGACAUUGAAGCGAAAGC-----UAAGA---- ....((((((.((((((.........)--)).))).))))))(((....))-----)....---- ( -14.10, z-score = -2.06, R) >consensus CAAAUCGAUGACAGACUAAACGAGCC___GAGUUGACAUUGAAGCGAAAGC_AGCUCAAGA____ ....((((((...((((.............))))..)))))).((....)).............. ( -8.53 = -8.54 + 0.01)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:31:17 2011