
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 16,420,152 – 16,420,246 |
| Length | 94 |
| Max. P | 0.690262 |

| Location | 16,420,152 – 16,420,246 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 71.52 |
| Shannon entropy | 0.51091 |
| G+C content | 0.51416 |
| Mean single sequence MFE | -27.57 |
| Consensus MFE | -7.32 |
| Energy contribution | -9.06 |
| Covariance contribution | 1.74 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.27 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.690262 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 16420152 94 - 24543557 ---------ACGCCAGA-AAGCUGGCAGUAGACAAACAACACUUUGUAACACCUC--GACAAGACACCUUU-----------UUCCUUCGAGGAGCAAACAUUUGCCAGCCG-GCCAG ---------..(((...-..((((((((...(((((......)))))....((((--((...((.......-----------.))..)))))).........)))))))).)-))... ( -26.20, z-score = -3.18, R) >droSim1.chr3L 15764129 94 - 22553184 ---------ACGCCAGA-AAGCUGGCAGUAGACAAACAACACUUUGUAACACCUC--GACAAGACACCUUU-----------UUCCUCCGAGGAGCAAACAUUCGCCAGCCG-GCCAG ---------..(((...-..((((((.((..(((((......))))).)).((((--(.............-----------......)))))...........)))))).)-))... ( -23.51, z-score = -2.37, R) >droSec1.super_0 8514427 94 - 21120651 ---------ACGCCAGA-AAGCUGGCAGUAGACAAACAACACUUUGUAACACCUC--GACAAGACACCUUU-----------UUCCUCCGAGGAGCAAACAUUCGCCAGCCG-GCCAG ---------..(((...-..((((((.((..(((((......))))).)).((((--(.............-----------......)))))...........)))))).)-))... ( -23.51, z-score = -2.37, R) >droYak2.chr3L 6561146 92 + 24197627 ---------AUGCCAGA-AAGCUGGCAGUAAACAAACAACACU--GUAACACCUC--GACAAGACACCUCU-----------UUCCUCCGAGGAGCAAACAUUUGCCAGCCG-GCCAG ---------..(((...-..((((((((.......(((....)--))....((((--(..((((....)))-----------).....))))).........)))))))).)-))... ( -23.30, z-score = -2.20, R) >droEre2.scaffold_4784 8372553 94 + 25762168 ---------ACGCCAGA-AAGCUGGCAGUAGACAAACAACACUUUGUAACACCUC--GACAAGACACCUUC-----------UGCCUCCAAGGAGCAAAUAUUUGCCAGCUG-GCCAG ---------..((((..-..((((((((((((...((((....))))......((--.....)).....))-----------)))(((....)))........)))))))))-))... ( -24.60, z-score = -1.86, R) >droAna3.scaffold_13337 9138948 116 + 23293914 CCGCCAGACGUGGCAGACGAGGUGGCAGAAGACAAACAACACUCUGCAACGCCUC--GACACGACAGGCUCAGACCCAGAAGAUCUUCUGGGGAGCAAACAUUUGCCAAAAGAGCCAA ..((((....))))...((((((((((((.............)))))..))))))--)........(((((...(((((((....)))))))(.((((....)))))....))))).. ( -46.22, z-score = -4.98, R) >droVir3.scaffold_13049 1414307 83 - 25233164 ------------------------GCAGCAGCCAACAGGCACUUGGUCUGGGCUCAUGGCAAGUCAACGAG---------CUGCAAAACAAAUAGCAAAUAUUUG-CAGCUG-ACAAG ------------------------((...((((..(((((.....)))))))))....))..((((....(---------(((((((..............))))-))))))-))... ( -25.64, z-score = -1.43, R) >consensus _________ACGCCAGA_AAGCUGGCAGUAGACAAACAACACUUUGUAACACCUC__GACAAGACACCUUU___________UUCCUCCGAGGAGCAAACAUUUGCCAGCCG_GCCAG ....................((((((((..............(((((...........)))))......................(((....))).......))))))))........ ( -7.32 = -9.06 + 1.74)
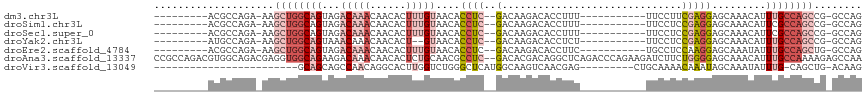

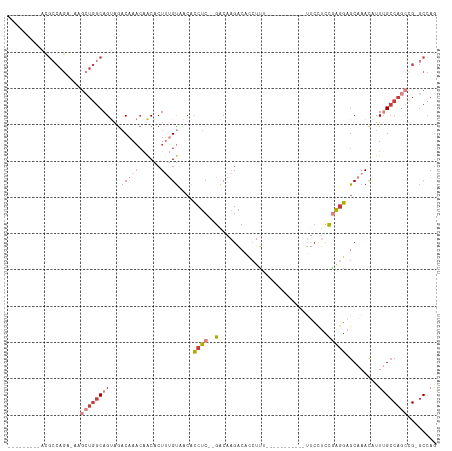
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:31:16 2011