
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 16,292,459 – 16,292,518 |
| Length | 59 |
| Max. P | 0.755064 |

| Location | 16,292,459 – 16,292,518 |
|---|---|
| Length | 59 |
| Sequences | 11 |
| Columns | 64 |
| Reading direction | reverse |
| Mean pairwise identity | 84.42 |
| Shannon entropy | 0.32877 |
| G+C content | 0.29969 |
| Mean single sequence MFE | -10.74 |
| Consensus MFE | -8.32 |
| Energy contribution | -8.12 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.755064 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 16292459 59 - 24543557 AAUUUACGUACAUGAAUUUAAUGUUUUAGCUGUG-----GCUAUCAUUAAGUCUAAUUGACUUU .................((((((...((((....-----)))).))))))(((.....)))... ( -10.20, z-score = -1.60, R) >droSim1.chr3L 15628391 59 - 22553184 AAUUUACGUACAUGAAUUUAAUGUUUUAGCUGUG-----GCUAUCAUUAAGUCUAAUUGACUUU .................((((((...((((....-----)))).))))))(((.....)))... ( -10.20, z-score = -1.60, R) >droSec1.super_0 8386491 59 - 21120651 AAUUUACGUACAUGAAUUUAAUGUUUUAGCUGUG-----GCUAUCAUUAAGUCUAAUUGACUUU .................((((((...((((....-----)))).))))))(((.....)))... ( -10.20, z-score = -1.60, R) >droYak2.chr3L 2294185 59 + 24197627 AAUUUACGUACAUGAAUUUAAUGUUUUAGCUGUG-----GCUAUCAUUAAGUCUAAUUGACUUU .................((((((...((((....-----)))).))))))(((.....)))... ( -10.20, z-score = -1.60, R) >droEre2.scaffold_4784 18535139 59 - 25762168 AAUUUACGUACAUGAAUUUAAUGUGCUAGCUGUG-----GCUAUCAUUAAGUCUAAUUGACUUU .......((((((.......))))))((((....-----)))).....(((((.....))))). ( -12.90, z-score = -2.21, R) >droAna3.scaffold_13337 12957305 56 + 23293914 --UUCUUGUACAUGAAUUUAAUGU-UUAGCUGGG-----GCUAUCAUUAAGUCUAAUUGACUUU --...............((((((.-.((((....-----)))).))))))(((.....)))... ( -10.20, z-score = -1.40, R) >dp4.chrXR_group8 1979952 63 - 9212921 AAUUUACGUACAUGAAUUUAAUGU-UUAGCUGCGGCUAUGCUAUCAUUAAGUCUAAUUGACUUU .................((((((.-.((((.........)))).))))))(((.....)))... ( -9.60, z-score = -0.52, R) >droPer1.super_24 480115 63 + 1556852 AAUUUACGUACAUGAAUUUAAUGU-UUAGCUGCGGCUAUGCUAUCAUUAAGUCUAAUUGACUUU .................((((((.-.((((.........)))).))))))(((.....)))... ( -9.60, z-score = -0.52, R) >droVir3.scaffold_13049 1264902 62 - 25233164 AGCGUAC-UACACGAAUUUAAUGU-GUAGUUUAGCCGAAGCUAUCAUUAAGUCUAAUUGACUUU .((..((-((((((.......)))-)))))...)).............(((((.....))))). ( -14.40, z-score = -2.28, R) >droMoj3.scaffold_6654 9801 62 + 2564135 AGCGUAC-CAAACAAAUUUAAUGU-GUAGUUUAGCCGAAGCUAUCAUUAAGUCUAAUUGACUUU .......-.........((((((.-(((((((.....)))))))))))))(((.....)))... ( -10.70, z-score = -1.57, R) >droGri2.scaffold_15110 2395475 62 - 24565398 AGCGUAC-AACACGAAUUUAAUGU-GUAGUUUAGCCACAGCUAUCAUUAAGUCUAAUUGACUUU .((..((-.(((((.......)))-)).))...)).............(((((.....))))). ( -9.90, z-score = -0.72, R) >consensus AAUUUACGUACAUGAAUUUAAUGU_UUAGCUGUG_____GCUAUCAUUAAGUCUAAUUGACUUU .................((((((...((((.........)))).))))))(((.....)))... ( -8.32 = -8.12 + -0.20)

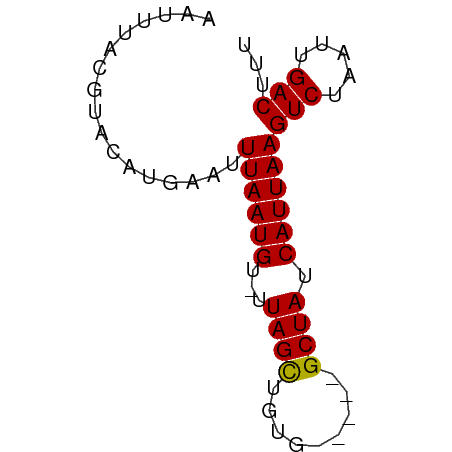
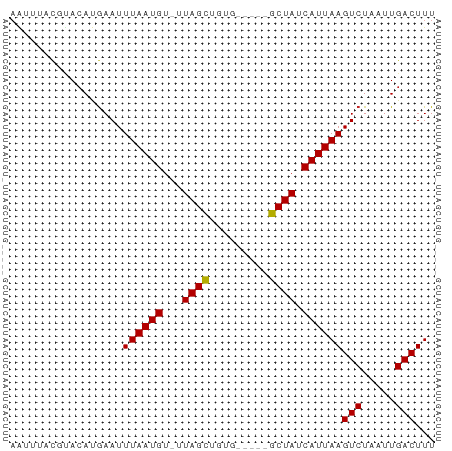
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:31:04 2011