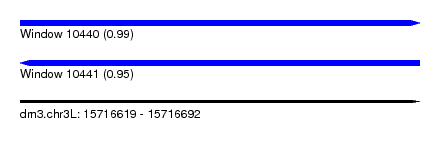
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,716,619 – 15,716,692 |
| Length | 73 |
| Max. P | 0.992979 |

| Location | 15,716,619 – 15,716,692 |
|---|---|
| Length | 73 |
| Sequences | 7 |
| Columns | 76 |
| Reading direction | forward |
| Mean pairwise identity | 71.57 |
| Shannon entropy | 0.58768 |
| G+C content | 0.41132 |
| Mean single sequence MFE | -18.69 |
| Consensus MFE | -9.36 |
| Energy contribution | -11.91 |
| Covariance contribution | 2.56 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.58 |
| SVM RNA-class probability | 0.992979 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 15716619 73 + 24543557 ---AAUGGGUGCUUCCCAAUCAGGCAUCAAAGUUUUGACUCUGUCAUUACUUUGUUUGUCCAUGUCAAAGGACAUU ---..((((.....))))....((((.((((((..((((...))))..))))))..)))).(((((....))))). ( -22.20, z-score = -2.97, R) >droEre2.scaffold_4784 17972708 73 + 25762168 ---AGUGGGUGCUUCCCAAUCAGGCAUCAAAGUUUUGACUCUAUCAUUACUUUGUUUGGCCAUGUCAAAGGACAUU ---..((((.....))))....(((..((((((..(((.....)))..))))))....)))(((((....))))). ( -22.40, z-score = -2.70, R) >droYak2.chr3L 18267500 76 + 24197627 AGUAGGGGGUGCUUCCCAAUCAGGCAUCAAAGUUUUGACUCCGUCAUUACUUUGUUUGUCCAUGUCAAAGGACAUU .....((((....)))).....((((.((((((..((((...))))..))))))..)))).(((((....))))). ( -22.30, z-score = -2.33, R) >droSec1.super_0 7827698 73 + 21120651 ---AGUGGGUGCUUCCCAAUCAGGCAUCAAAGUUUUGACUCUGUCAUUACUUUGUUUGUCCAUGUCAAAGGACAUU ---..((((.....))))....((((.((((((..((((...))))..))))))..)))).(((((....))))). ( -22.10, z-score = -2.68, R) >droSim1.chr3L 15055606 73 + 22553184 ---AAUGGGUGCUUCCCAAUCAGGCAUCAAAGUUUUGACUCUGUCAUUACUUUGUUUGUCCAUGUCAAAGGACAUU ---..((((.....))))....((((.((((((..((((...))))..))))))..)))).(((((....))))). ( -22.20, z-score = -2.97, R) >droAna3.scaffold_13337 11855448 56 - 23293914 ---AUAGAGUUCUCCCCAAUCAGGCAUCA-----------------UUACAUUGUUUGCCCCUGUCAAAGGACAUU ---.....(((((.........((((.((-----------------......))..))))........)))))... ( -9.33, z-score = -0.72, R) >droWil1.scaffold_180727 1122631 55 - 2741493 ------------------ACUAAAAACUUUUGUCUUGUCUGCGACAAUUGUUUGCUU---CAAGUCCGACGAGAAC ------------------..............(((((((.(.(((..(((.......---)))))))))))))).. ( -10.30, z-score = -0.82, R) >consensus ___AGUGGGUGCUUCCCAAUCAGGCAUCAAAGUUUUGACUCUGUCAUUACUUUGUUUGUCCAUGUCAAAGGACAUU ......(((.....))).....((((.((((((..((((...))))..))))))..)))).(((((....))))). ( -9.36 = -11.91 + 2.56)
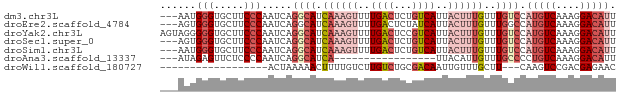
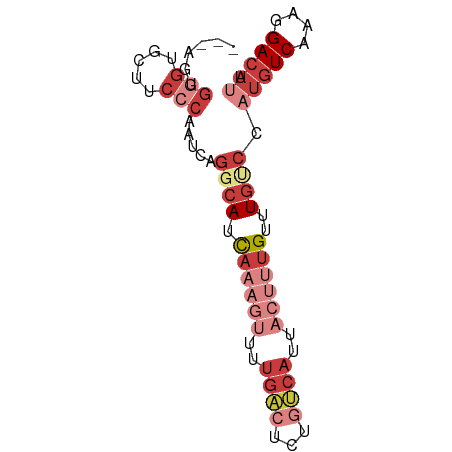
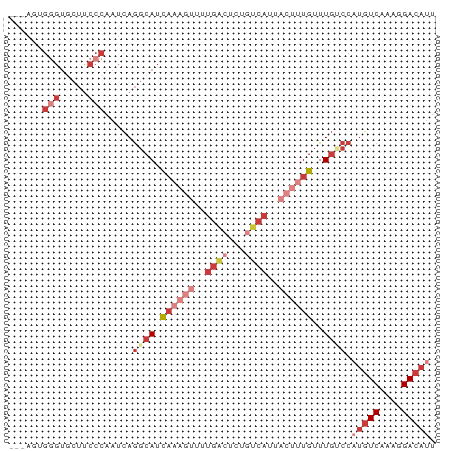
| Location | 15,716,619 – 15,716,692 |
|---|---|
| Length | 73 |
| Sequences | 7 |
| Columns | 76 |
| Reading direction | reverse |
| Mean pairwise identity | 71.57 |
| Shannon entropy | 0.58768 |
| G+C content | 0.41132 |
| Mean single sequence MFE | -16.19 |
| Consensus MFE | -8.81 |
| Energy contribution | -10.71 |
| Covariance contribution | 1.91 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.948534 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 15716619 73 - 24543557 AAUGUCCUUUGACAUGGACAAACAAAGUAAUGACAGAGUCAAAACUUUGAUGCCUGAUUGGGAAGCACCCAUU--- ..(((((........)))))..((((((..((((...))))..)))))).........((((.....))))..--- ( -19.50, z-score = -2.40, R) >droEre2.scaffold_4784 17972708 73 - 25762168 AAUGUCCUUUGACAUGGCCAAACAAAGUAAUGAUAGAGUCAAAACUUUGAUGCCUGAUUGGGAAGCACCCACU--- .(((((....)))))(((....((((((..((((...))))..))))))..)))....((((.....))))..--- ( -17.90, z-score = -1.77, R) >droYak2.chr3L 18267500 76 - 24197627 AAUGUCCUUUGACAUGGACAAACAAAGUAAUGACGGAGUCAAAACUUUGAUGCCUGAUUGGGAAGCACCCCCUACU ..(((((........)))))..((((((..((((...))))..))))))..........(((......)))..... ( -18.50, z-score = -1.76, R) >droSec1.super_0 7827698 73 - 21120651 AAUGUCCUUUGACAUGGACAAACAAAGUAAUGACAGAGUCAAAACUUUGAUGCCUGAUUGGGAAGCACCCACU--- ..(((((........)))))..((((((..((((...))))..)))))).........((((.....))))..--- ( -19.40, z-score = -2.33, R) >droSim1.chr3L 15055606 73 - 22553184 AAUGUCCUUUGACAUGGACAAACAAAGUAAUGACAGAGUCAAAACUUUGAUGCCUGAUUGGGAAGCACCCAUU--- ..(((((........)))))..((((((..((((...))))..)))))).........((((.....))))..--- ( -19.50, z-score = -2.40, R) >droAna3.scaffold_13337 11855448 56 + 23293914 AAUGUCCUUUGACAGGGGCAAACAAUGUAA-----------------UGAUGCCUGAUUGGGGAGAACUCUAU--- ..(.(((((...((((.(((..........-----------------)).).))))...))))).).......--- ( -10.40, z-score = 0.15, R) >droWil1.scaffold_180727 1122631 55 + 2741493 GUUCUCGUCGGACUUG---AAGCAAACAAUUGUCGCAGACAAGACAAAAGUUUUUAGU------------------ ((((.....))))...---....((((..(((((........)))))..)))).....------------------ ( -8.10, z-score = 0.48, R) >consensus AAUGUCCUUUGACAUGGACAAACAAAGUAAUGACAGAGUCAAAACUUUGAUGCCUGAUUGGGAAGCACCCACU___ ..(((((........)))))..((((((..((((...))))..))))))..........(((.....)))...... ( -8.81 = -10.71 + 1.91)

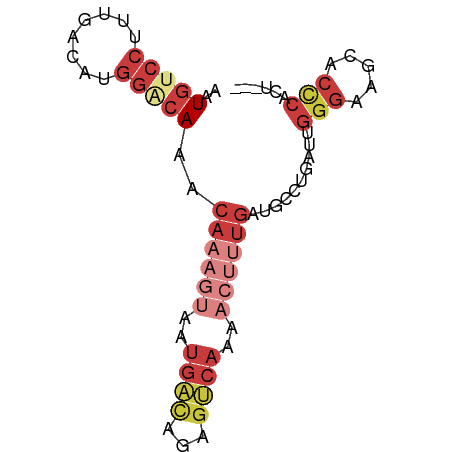
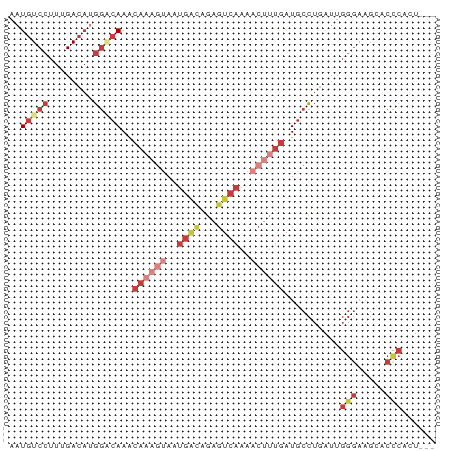
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:29:40 2011