
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 5,347,803 – 5,347,894 |
| Length | 91 |
| Max. P | 0.528865 |

| Location | 5,347,803 – 5,347,894 |
|---|---|
| Length | 91 |
| Sequences | 10 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 71.55 |
| Shannon entropy | 0.55222 |
| G+C content | 0.41227 |
| Mean single sequence MFE | -11.12 |
| Consensus MFE | -5.71 |
| Energy contribution | -5.52 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.18 |
| Mean z-score | -0.99 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.528865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 5347803 91 + 23011544 GUAUACGCUUUGGGGUUAG--CCCCCCGUC---CGACCGAUAAACUCAUAAAACAUCGAA-GAAUUGAAGCGCUUCGAUUUCAAUUUACCAUAAACG .....((...(((((....--..)))))..---))..((((.............))))..-(((((((((((...)).))))))))).......... ( -15.52, z-score = -0.26, R) >droSim1.chr2L 5140139 96 + 22036055 GACUACCCUUUGGGUGUAGCCCCCCACCUCGUCCGACCGAUAAACUCAUAAAACAUCGAA-GAAUUGAAGCGCUUCGAUUUCAAUUUAGCAUAAACG (.(((..(((((((((........))))(((......)))................))))-)((((((((((...)).))))))))))))....... ( -13.30, z-score = 0.19, R) >droSec1.super_5 3424255 92 + 5866729 GACUACCCUUUGGGUGUAGC-CCCCCCAUC---CGACCGAUAAACUCAUAAAACAUCGAA-GAAUUGAAACGCUUCGAUUUCAAUUUACCAUAAACG .........(((((((....-.....))))---))).((((.............))))..-(((((((((((...)).))))))))).......... ( -12.72, z-score = -0.93, R) >droYak2.chr2L 8472895 90 - 22324452 GACUACCCCCCCUUCGUG---UGCCCCGUC---UGACCGAUAAACUCAUAAAACAUCGAA-GAAUUGAAGCGCUUCGAUUUCAAUUUACCAUAAACG ...........(((((((---(........---...................))).))))-)((((((((((...)).))))))))........... ( -10.03, z-score = -0.72, R) >droEre2.scaffold_4929 5428693 88 + 26641161 GACUACCCUCUGGGAG-----CCCCGUCUC---CGACCGAUAAACUCAUAAAACAUCGAA-GAAUUGAAGCGCUUCGAUUUCAAUUUACCAUAAACG ............((((-----......)))---)...((((.............))))..-(((((((((((...)).))))))))).......... ( -11.52, z-score = 0.14, R) >droAna3.scaffold_12916 6370997 70 - 16180835 -----------------------GACUACC---UGCCCAAUAAACUCAUAAAACAUCGAA-CAAUUGAAGCGCUUCGAUUUCAAUUUACCAUAAACG -----------------------.......---........................(((-..((((((....)))))))))............... ( -4.20, z-score = -0.82, R) >dp4.chr4_group1 4627358 75 + 5278887 ------------------GACUACCCAACC---GAACCGAUAAACUCAUAAAACAUUGGGGAAAUUGAAGCGCUUCGAUUUCAAUUUACCAUAAAA- ------------------.....(((((..---.....((.....))........)))))(((((((((....)))))))))..............- ( -12.96, z-score = -2.31, R) >droPer1.super_5 6229133 75 - 6813705 ------------------GACUACCCAACC---GAACCGAUAAACUCAUAAAACAUUGGGGAAAUUGAAGCGCCUCGAUUUCAAUUUACCAUAAAA- ------------------.....(((((..---.....((.....))........)))))((((((((......))))))))..............- ( -11.56, z-score = -1.65, R) >droVir3.scaffold_12963 257030 69 - 20206255 ------------------GACUACCC--CC---GC-GGAAUAAACUCAUAAAGCAUCG---AAAUUGAAGCGCUUCGAUUUCAAUUUACCAUAAAA- ------------------........--..---((-.((......)).....))...(---((((((((....)))))))))..............- ( -10.00, z-score = -1.45, R) >droGri2.scaffold_15252 5769503 70 - 17193109 ------------------GACUACCC--CC---AAAGGAAUAAACUCAUAAAGCAUCG---AAAUUGAAGCGCUUCGAUUUCAAUUUACCAUAAAA- ------------------.....((.--..---...))...................(---((((((((....)))))))))..............- ( -9.40, z-score = -2.06, R) >consensus __________________G__UACCCCACC___CGACCGAUAAACUCAUAAAACAUCGAA_GAAUUGAAGCGCUUCGAUUUCAAUUUACCAUAAACG .............................................................(((((((((((...)).))))))))).......... ( -5.71 = -5.52 + -0.19)


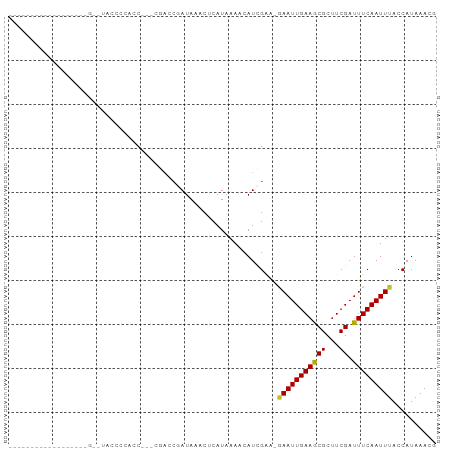
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:18:47 2011