| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,538,096 – 15,538,183 |
| Length | 87 |
| Max. P | 0.930471 |

| Location | 15,538,096 – 15,538,183 |
|---|---|
| Length | 87 |
| Sequences | 9 |
| Columns | 88 |
| Reading direction | reverse |
| Mean pairwise identity | 83.46 |
| Shannon entropy | 0.33868 |
| G+C content | 0.40274 |
| Mean single sequence MFE | -15.88 |
| Consensus MFE | -10.71 |
| Energy contribution | -10.64 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.35 |
| SVM RNA-class probability | 0.930471 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 15538096 87 - 24543557 CAACGAACCUC-GGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGCUACCAAAACGGGCAACACAGAUACAGACAAACACACGUAC ..(((....((-((((((((((((...))))))))))))))....(((..((.....))))).....................))).. ( -18.00, z-score = -2.82, R) >droSim1.chr3L 14879597 87 - 22553184 CAACGAACCUC-GGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGCUACCAAAACGGGCAACACAGAUACAGACAAACACACGUAC ..(((....((-((((((((((((...))))))))))))))....(((..((.....))))).....................))).. ( -18.00, z-score = -2.82, R) >droSec1.super_0 7652011 87 - 21120651 CAACGAACCUC-GGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGCUACCAAAACGGGCAACACAGAUAGAGACAAACACACGUAC ..(((....((-((((((((((((...))))))))))))))....(((..((.....))))).....................))).. ( -18.00, z-score = -2.67, R) >droYak2.chr3L 18054085 85 - 24197627 CAACGAACCUU-GGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGCUACCAAAACGGGCAACACAGAUACACACCAACACGUAC-- ..(((....(.-.(((((((((((...)))))))))))..)....(((..((.....)))))...................)))..-- ( -16.60, z-score = -2.10, R) >droEre2.scaffold_4784 17795817 85 - 25762168 CAACGAACCUU-GGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGCUACCAAAACGGGCAACACAGAUACACACAAACACGUAC-- ..(((....(.-.(((((((((((...)))))))))))..)....(((..((.....)))))...................)))..-- ( -16.60, z-score = -2.37, R) >droAna3.scaffold_13337 6487265 70 - 23293914 CAAUGAACCCCGGGGAAUUGUCGAGAAUUAGCAAUUCUUGAAAAAUGCUACCAAAACGGGCAACACACAC------------------ ........((((.((.(((.((((((((.....))))))))..)))....))....))))..........------------------ ( -14.10, z-score = -1.28, R) >droWil1.scaffold_181136 1874725 70 - 2313701 CAAUGAACUU--UGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGCUACCAAAACGGGCAACAGACACAC---------------- ........((--((((((((((((...))))))))))........(((..((.....))))).)))).....---------------- ( -11.60, z-score = -1.31, R) >droMoj3.scaffold_6680 8027722 86 - 24764193 CAACGAACCU--UGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGUUACCAAAACGGGCAACAUAAACACACACAGACAGAGACAC ........((--((((((((((((...)))))))))))......(((((.((......)).)))))...............))).... ( -14.20, z-score = -1.66, R) >droGri2.scaffold_15110 7979154 83 + 24565398 CAACGAACCU--UGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGUUACCAAAACGGGCAACAUAAACACACACAAGCACAUG--- ........((--((((((((((((...)))))))))).......(((((.((......)).))))).........))))......--- ( -15.80, z-score = -2.17, R) >consensus CAACGAACCUC_GGGAAUUGUUGAGAAUUAGCAAUUCUUGAAAAAUGCUACCAAAACGGGCAACACAGAUACACACAAACACACG___ .............(((((((((((...))))))))))).......(((..((.....))))).......................... (-10.71 = -10.64 + -0.06)
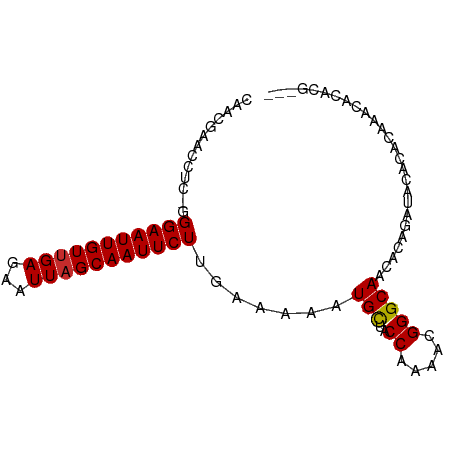


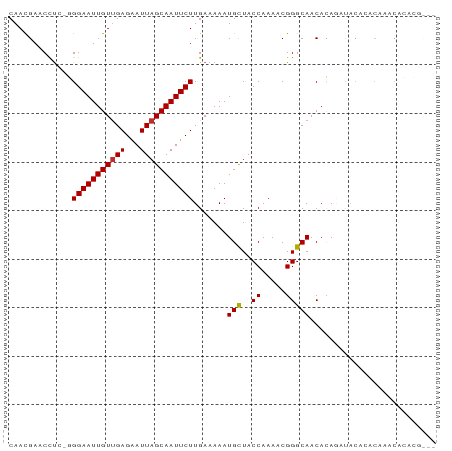
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:29:10 2011