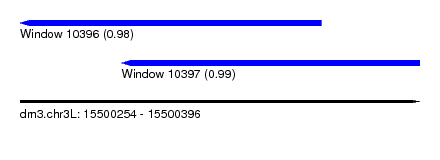

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,500,254 – 15,500,396 |
| Length | 142 |
| Max. P | 0.987849 |

| Location | 15,500,254 – 15,500,361 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 85.89 |
| Shannon entropy | 0.24019 |
| G+C content | 0.48696 |
| Mean single sequence MFE | -33.12 |
| Consensus MFE | -27.10 |
| Energy contribution | -27.86 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.977004 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
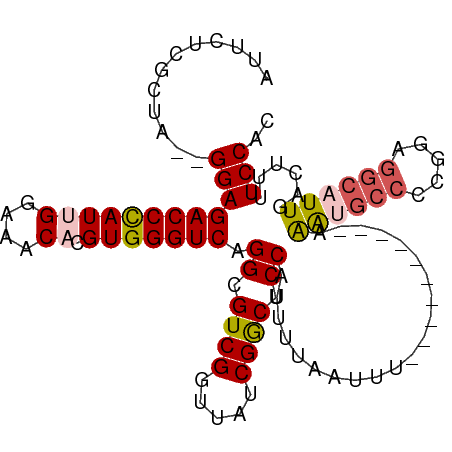
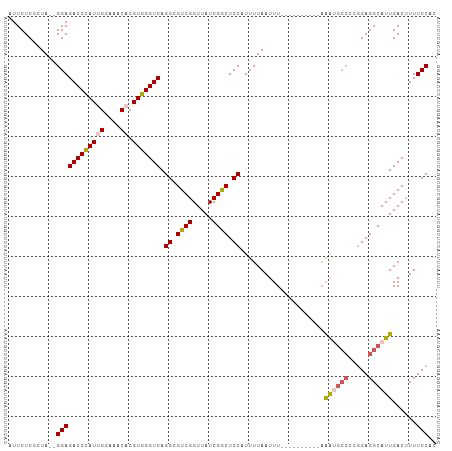
>dm3.chr3L 15500254 107 - 24543557 AUUCUCGCUAGGGGAGACCCAUUGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUUAAAUGCCCAUUGAGGCCAUUGAGGCAUUGACUUUUCCAC ..........((..((((((((((....)).))))))).((.((((.....)))).))...........((((((((.((....)).)).))))))....)..)).. ( -36.40, z-score = -2.31, R) >droEre2.scaffold_4784 17756202 89 - 25762168 AUUCUCCCUA--GGAGACCCAUUGGAAACACGUGGGUCAGGCGUCGGUUAUCGACUCCUUUUUAAUUU----------AAG-----CGGAGGCAUUGAC-UUUCCAC ..(((((((.--.(((((((((((....)).)))))))(((.((((.....)))).)))......)).----------.))-----.))))).......-....... ( -30.60, z-score = -2.87, R) >droYak2.chr3L 18015551 95 - 24197627 AUUCUCCCUA--GGAGACCCAUUGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUU----------AAAUGCCCCGGAGGCAUUGACUUUUCCAC ..........--((((((((((((....)).))))))).((.((((.....)))).))..........----------.((((((.....))))))......))).. ( -32.30, z-score = -2.35, R) >droSec1.super_0 7609600 105 - 21120651 AUUCUCGCUA--GGAGACCCAUGGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUUAAAUGCAUUUAAAUGCCCCGGAGGCAUUGACUUUUCCAC ..........--((((((((((((....).)))))))).((.((((.....)))).)).....................((((((.....))))))......))).. ( -34.30, z-score = -2.33, R) >droSim1.chr3L 14841073 95 - 22553184 AUUCUCGCUA--GGAGACCUAUGGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUU----------AAAUGCCCCGGAGGCAUUGACUUUUCCAC ..........--((((((((((((....).)))))))).((.((((.....)))).))..........----------.((((((.....))))))......))).. ( -32.00, z-score = -2.22, R) >consensus AUUCUCGCUA__GGAGACCCAUUGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUU__________AAAUGCCCCGGAGGCAUUGACUUUUCCAC ............((((((((((((....)).))))))).((.((((.....)))).)).....................((((((.....))))))......))).. (-27.10 = -27.86 + 0.76)

| Location | 15,500,290 – 15,500,396 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 95.04 |
| Shannon entropy | 0.08431 |
| G+C content | 0.45784 |
| Mean single sequence MFE | -33.30 |
| Consensus MFE | -30.82 |
| Energy contribution | -31.14 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.93 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.30 |
| SVM RNA-class probability | 0.987849 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
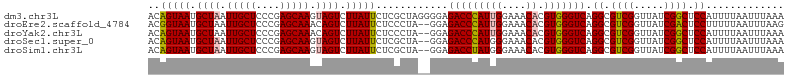
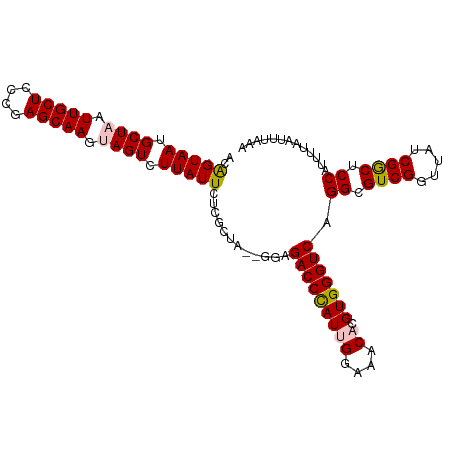
>dm3.chr3L 15500290 106 - 24543557 ACAGUAAUGCUAAUUGCUCCCGAGCAAGUAGUCUUAUUCUCGCUAGGGGAGACCCAUUGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUUAAA ....(((.((((.(((((....))))).)))).)))(((((....)))))(((((((((....)).))))))).((.((((.....)))).))............. ( -35.20, z-score = -2.77, R) >droEre2.scaffold_4784 17756222 104 - 25762168 ACGGUAAUGCUAAUUGCUCCCGAGCAAACAGUCUUAUUCUCCCUA--GGAGACCCAUUGGAAACACGUGGGUCAGGCGUCGGUUAUCGACUCCUUUUUAAUUUAAG ..(((((.(((..(((((....)))))..))).))))).(((...--)))(((((((((....)).)))))))(((.((((.....)))).)))............ ( -31.80, z-score = -2.57, R) >droYak2.chr3L 18015577 104 - 24197627 ACAGUAAUGCUAAUUGCUCCCGAGCAAACAGUCUUAUUCUCCCUA--GGAGACCCAUUGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUUAAA ..(((((.(((..(((((....)))))..))).))))).(((...--)))(((((((((....)).))))))).((.((((.....)))).))............. ( -31.20, z-score = -2.44, R) >droSec1.super_0 7609636 104 - 21120651 ACAGUAAUGCUAAUUGCUCCCGAGCAAGUAGUCUUAUUCUCGCUA--GGAGACCCAUGGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUUAAA ..(((((.((((.(((((....))))).)))).))))).....((--((((((((((((....).)))))))).((.((((.....)))).)).)))))....... ( -35.30, z-score = -3.15, R) >droSim1.chr3L 14841099 104 - 22553184 ACAGUAAUGCUAAUUGCUCCCGAGCAAGUAGUCUUAUUCUCGCUA--GGAGACCUAUGGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUUAAA ..(((((.((((.(((((....))))).)))).))))).....((--((((((((((((....).)))))))).((.((((.....)))).)).)))))....... ( -33.00, z-score = -2.46, R) >consensus ACAGUAAUGCUAAUUGCUCCCGAGCAAGUAGUCUUAUUCUCGCUA__GGAGACCCAUUGGAAACACGUGGGUCAGGCGUCGGUUAUCGGCUCCAUUUUAAUUUAAA ..(((((.((((.(((((....))))).)))).)))))............(((((((((....)).))))))).((.((((.....)))).))............. (-30.82 = -31.14 + 0.32)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:29:03 2011