
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,384,696 – 15,384,791 |
| Length | 95 |
| Max. P | 0.718879 |

| Location | 15,384,696 – 15,384,791 |
|---|---|
| Length | 95 |
| Sequences | 9 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 75.19 |
| Shannon entropy | 0.51609 |
| G+C content | 0.50300 |
| Mean single sequence MFE | -26.26 |
| Consensus MFE | -13.05 |
| Energy contribution | -13.33 |
| Covariance contribution | 0.29 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.718879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 15384696 95 + 24543557 -CAAAGAGGGGAGGAAAA---------AGGGGUCCCGGGGCUUCGGGCCACAUUCUAUGCCACAGAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCAGGCAG -...((((((((......---------.....)))).((.(....).))...)))).((((...(((((((.....))))))).....(((......))))))). ( -24.80, z-score = -0.75, R) >droSim1.chr3L 14724794 104 + 22553184 -CAAAGAUGAGGCGGGAAGGGGAAAAAAGGGGUCCCGGGGCUUCGGGCCACACUCUAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCAGGCAG -.......(((.(((((...............)))))((.(....).))...)))..((((...(((((((.....))))))).....(((......))))))). ( -27.66, z-score = -0.82, R) >droSec1.super_0 7493963 103 + 21120651 -CAAAGAGGAGGCGGGAAGAGGAAAAAAGGGGUCC-GGGGCUUCGGGCCACACUCUAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCAGGCAG -...(((((.(((..((((.(((.........)))-....))))..))).).)))).((((...(((((((.....))))))).....(((......))))))). ( -28.20, z-score = -1.91, R) >droYak2.chr3L 17900789 95 + 24197627 -CAAAGAAGAGGCGGGAA---------AGGGGUCCCGGGGCUUCGGGCCACACUCUAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCAGGCAG -......((((.(((((.---------.....)))))((.(....).))...)))).((((...(((((((.....))))))).....(((......))))))). ( -30.60, z-score = -2.36, R) >droEre2.scaffold_4784 17643203 96 + 25762168 -CAAAGAAGAGGCGGGAA--------AAGGGGUCCCGUGGCUUCGGGCCACACUCUAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCAGGCAG -......((((..((((.--------......))))(((((.....))))).)))).((((...(((((((.....))))))).....(((......))))))). ( -31.10, z-score = -2.51, R) >droAna3.scaffold_13337 5814886 102 - 23293914 CAGAAAAGGGGAGGCUACUGGGUGGCAAGAUGGCUAGGGGG---GCGCCACACUCAAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCAGGCAA ..........((((((((...)))))....((((.......---..))))..)))..((((...(((((((.....))))))).....(((......))))))). ( -24.40, z-score = -0.17, R) >droPer1.super_12 1364620 83 - 2414086 -------------CAGAA---------AGGGGACUAGGGGGCUAUGACCACAGUCAAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAACGCGCACAAGCGGGCAU -------------.....---------....((((..((........))..)))).(((((...(((((((.....))))))).....(((......)))))))) ( -21.30, z-score = -2.54, R) >droVir3.scaffold_13049 20165170 91 + 25233164 -AGGAAAGGCGGCAGCAC---------AGG-GAUAACGGCG---GGCCAGCACUCAAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCGGGCAA -..((...(((((.((..---------.(.-.....).)).---.))).))..))..((((...(((((((.....))))))).......(((....))))))). ( -22.70, z-score = -0.75, R) >droMoj3.scaffold_6680 7865364 93 + 24764193 CGGAAGAGGGCGCUGCAC---------AGGCAAUGCCAACG---GGGCGGUACUCAAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCAGGCAG (....).(((..((((.(---------.(((...)))....---).))))..)))..((((...(((((((.....))))))).....(((......))))))). ( -25.60, z-score = -0.97, R) >consensus _CAAAGAGGAGGCGGGAA_________AGGGGUCCCGGGGCUUCGGGCCACACUCUAUGCCACAAAAUUAUGAUAAAUGAUUUGGAAAUGCGCACAAGCAGGCAG ......................................(((.....)))........((((...(((((((.....))))))).....(((......))))))). (-13.05 = -13.33 + 0.29)


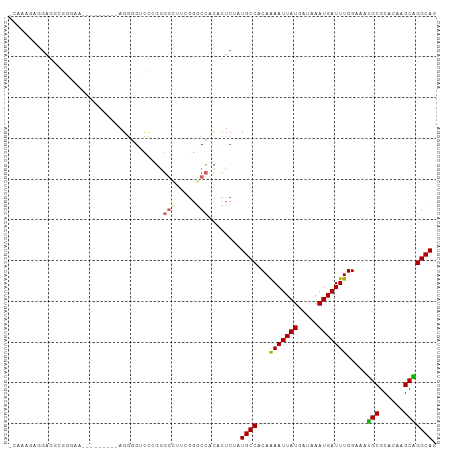
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:28:42 2011