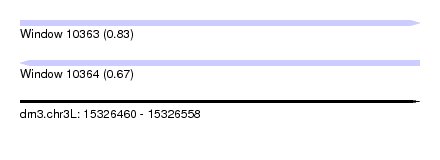

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,326,460 – 15,326,558 |
| Length | 98 |
| Max. P | 0.831266 |

| Location | 15,326,460 – 15,326,558 |
|---|---|
| Length | 98 |
| Sequences | 11 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 82.10 |
| Shannon entropy | 0.36304 |
| G+C content | 0.40605 |
| Mean single sequence MFE | -22.95 |
| Consensus MFE | -19.29 |
| Energy contribution | -18.99 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.831266 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
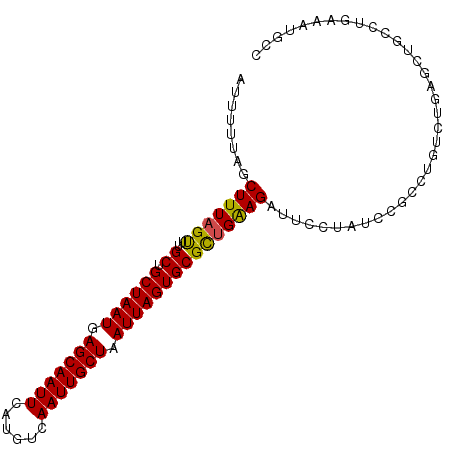
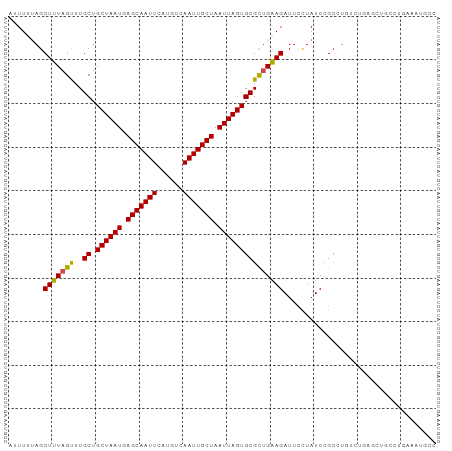
>dm3.chr3L 15326460 98 + 24543557 AUUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGGAGAUUCCUAUCCGCCUGUCUGAGCUGCCUGAAAUGCC ..((((((...((((((((.((((((.(((((((......))))))).))))))))((.(((........)))))......)))))).)))))).... ( -24.80, z-score = -1.19, R) >droSim1.chr3L 14664817 98 + 22553184 AUUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGGAGAUUCCUAUCCGCCUGUCUGAGCUGCCUGAAAUGCC ..((((((...((((((((.((((((.(((((((......))))))).))))))))((.(((........)))))......)))))).)))))).... ( -24.80, z-score = -1.19, R) >droSec1.super_0 7435590 98 + 21120651 AUUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGGAGAUUCCUAUCCGCCUGUCUGAGCUGCCUGAAAUGCC ..((((((...((((((((.((((((.(((((((......))))))).))))))))((.(((........)))))......)))))).)))))).... ( -24.80, z-score = -1.19, R) >droYak2.chr3L 17836988 98 + 24197627 AUUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGGAGAUUCCUAUCCGCCUGUCUGAGCUGCCUGAAAUGCC ..((((((...((((((((.((((((.(((((((......))))))).))))))))((.(((........)))))......)))))).)))))).... ( -24.80, z-score = -1.19, R) >droEre2.scaffold_4784 17584656 98 + 25762168 AUUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGGAGAUUCCUAUCCGCCUGUCUGAGCUGCCUGAAAUGCC ..((((((...((((((((.((((((.(((((((......))))))).))))))))((.(((........)))))......)))))).)))))).... ( -24.80, z-score = -1.19, R) >droAna3.scaffold_13337 5619250 98 - 23293914 AUUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGAAGAUUCCUAUCCGGCUGUCUGAGCUGCCCGAAAUGCC ........(((((((..((.((((((.(((((((......))))))).))))))))))))))).......((.(((.((....)).))).))...... ( -26.30, z-score = -1.70, R) >dp4.chrXR_group6 12886775 91 - 13314419 -AUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGUCGAAGAUUUCUAUCUGCCUGUCUGCCUCAGUGAC------ -.....(((........)))((((((.(((((((......))))))).))))))..((((.((((....))))..(((.......)))))))------ ( -18.30, z-score = 0.06, R) >droPer1.super_12 1305186 91 - 2414086 -AUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGUCGAAGAUUUCUAUCUGCCUGUCUGCCUCAGUGAC------ -.....(((........)))((((((.(((((((......))))))).))))))..((((.((((....))))..(((.......)))))))------ ( -18.30, z-score = 0.06, R) >droVir3.scaffold_13049 5124388 77 - 25233164 AGUUGUAGCUUUAGCUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGAAGAUUUUUAUCUGCUU--------------------- ....(((((((((((..((.((((((.(((((((......))))))).)))))))))))))))........))))..--------------------- ( -22.50, z-score = -1.76, R) >droMoj3.scaffold_6680 16953972 77 - 24764193 AGUUUCAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGAAGAUUUUUAUCUGCUU--------------------- ........(((((((..((.((((((.(((((((......))))))).)))))))))))))))..............--------------------- ( -20.30, z-score = -1.30, R) >droGri2.scaffold_15110 20054659 77 + 24565398 CGUUGAAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGAAGAUUUCUAUCUGCCU--------------------- ....(((((((((((..((.((((((.(((((((......))))))).))))))))))))))).)))).........--------------------- ( -22.80, z-score = -2.51, R) >consensus AUUUUUAGCUUUAGUUUGCUGCUAAUGAGCAAUUCAUGUCAAUUGCUAAUUAGUGCGCUGAAGAUUCCUAUCCGCCUGUCUGAGCUGCCUGAAAUGCC ........(((((((..((.((((((.(((((((......))))))).)))))))))))))))................................... (-19.29 = -18.99 + -0.30)

| Location | 15,326,460 – 15,326,558 |
|---|---|
| Length | 98 |
| Sequences | 11 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 82.10 |
| Shannon entropy | 0.36304 |
| G+C content | 0.40605 |
| Mean single sequence MFE | -20.65 |
| Consensus MFE | -16.25 |
| Energy contribution | -15.95 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.10 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.672591 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
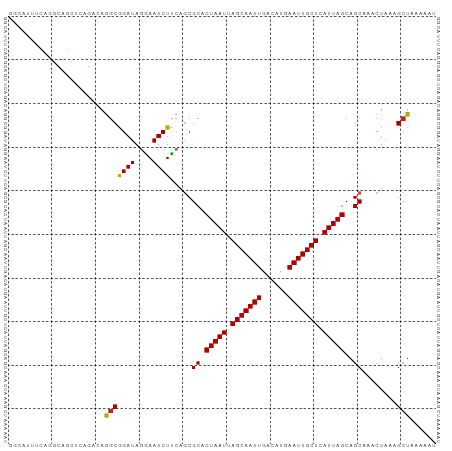
>dm3.chr3L 15326460 98 - 24543557 GGCAUUUCAGGCAGCUCAGACAGGCGGAUAGGAAUCUCCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAAU (((.......((.(((......)))(((........))).))((.(((((.(((((((......))))))).)))))..))........)))...... ( -22.80, z-score = -0.89, R) >droSim1.chr3L 14664817 98 - 22553184 GGCAUUUCAGGCAGCUCAGACAGGCGGAUAGGAAUCUCCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAAU (((.......((.(((......)))(((........))).))((.(((((.(((((((......))))))).)))))..))........)))...... ( -22.80, z-score = -0.89, R) >droSec1.super_0 7435590 98 - 21120651 GGCAUUUCAGGCAGCUCAGACAGGCGGAUAGGAAUCUCCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAAU (((.......((.(((......)))(((........))).))((.(((((.(((((((......))))))).)))))..))........)))...... ( -22.80, z-score = -0.89, R) >droYak2.chr3L 17836988 98 - 24197627 GGCAUUUCAGGCAGCUCAGACAGGCGGAUAGGAAUCUCCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAAU (((.......((.(((......)))(((........))).))((.(((((.(((((((......))))))).)))))..))........)))...... ( -22.80, z-score = -0.89, R) >droEre2.scaffold_4784 17584656 98 - 25762168 GGCAUUUCAGGCAGCUCAGACAGGCGGAUAGGAAUCUCCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAAU (((.......((.(((......)))(((........))).))((.(((((.(((((((......))))))).)))))..))........)))...... ( -22.80, z-score = -0.89, R) >droAna3.scaffold_13337 5619250 98 + 23293914 GGCAUUUCGGGCAGCUCAGACAGCCGGAUAGGAAUCUUCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAAU ...((((..(((.(((.(((...((.....))..)))..)))((.(((((.(((((((......))))))).)))))..))........)))..)))) ( -23.50, z-score = -1.09, R) >dp4.chrXR_group6 12886775 91 + 13314419 ------GUCACUGAGGCAGACAGGCAGAUAGAAAUCUUCGACGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAU- ------(((.(((.......)))(.((((....)))).))))((.(((((.(((((((......))))))).)))))..))................- ( -18.80, z-score = -0.92, R) >droPer1.super_12 1305186 91 + 2414086 ------GUCACUGAGGCAGACAGGCAGAUAGAAAUCUUCGACGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAU- ------(((.(((.......)))(.((((....)))).))))((.(((((.(((((((......))))))).)))))..))................- ( -18.80, z-score = -0.92, R) >droVir3.scaffold_13049 5124388 77 + 25233164 ---------------------AAGCAGAUAAAAAUCUUCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAGCUAAAGCUACAACU ---------------------.(((((((....))))..(((((.(((((.(((((((......))))))).)))))..))..)))...)))...... ( -17.70, z-score = -1.65, R) >droMoj3.scaffold_6680 16953972 77 + 24764193 ---------------------AAGCAGAUAAAAAUCUUCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUGAAACU ---------------------...............((((((((.(((((.(((((((......))))))).)))))..))........))))))... ( -17.40, z-score = -1.57, R) >droGri2.scaffold_15110 20054659 77 - 24565398 ---------------------AGGCAGAUAGAAAUCUUCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUUCAACG ---------------------((((((((....))))...((...(((((.(((((((......))))))).)))))..))........))))..... ( -17.00, z-score = -1.54, R) >consensus GGCAUUUCAGGCAGCUCAGACAGGCGGAUAGGAAUCUUCAGCGCACUAAUUAGCAAUUGACAUGAAUUGCUCAUUAGCAGCAAACUAAAGCUAAAAAU ......................(((((((....)))).....((.(((((.(((((((......))))))).)))))..))........)))...... (-16.25 = -15.95 + -0.30)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:28:35 2011