
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,280,936 – 15,281,035 |
| Length | 99 |
| Max. P | 0.519203 |

| Location | 15,280,936 – 15,281,035 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 88.86 |
| Shannon entropy | 0.22436 |
| G+C content | 0.36668 |
| Mean single sequence MFE | -23.87 |
| Consensus MFE | -20.01 |
| Energy contribution | -20.53 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.519203 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 15280936 99 - 24543557 -------------UUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUGAACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGUGCAAUGCAAC -------------.(((((.....((....))..)))))((((((...((((((.....))))))...((((((........))))))))))))...(((((.....))))) ( -23.00, z-score = -1.55, R) >droSim1.chr3L 14621589 99 - 22553184 -------------UUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUGAACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGUGCAAUGCAAC -------------.(((((.....((....))..)))))((((((...((((((.....))))))...((((((........))))))))))))...(((((.....))))) ( -23.00, z-score = -1.55, R) >droSec1.super_0 7392864 99 - 21120651 -------------CUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUGAACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGUGCAAUGCAAC -------------.(((((.....((....))..)))))((((((...((((((.....))))))...((((((........))))))))))))...(((((.....))))) ( -22.90, z-score = -1.51, R) >droEre2.scaffold_4784 17541664 108 - 25762168 ----CUUCCUGCUUUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUGAACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGUGCAAUGCAAC ----.....(((..((((......((....))..((((.((((((...((((((.....))))))...((((((........))))))))))))..))))..)))).))).. ( -23.60, z-score = -0.91, R) >droAna3.scaffold_13337 5578338 99 + 23293914 -------------UUUCCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUGAACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGUGCAAUGCACC -------------(((((......((....)).((((((..(((....((((((.....)))))).)))..))))))...........))))).......((((...)))). ( -20.00, z-score = -0.81, R) >droWil1.scaffold_180955 2186808 111 + 2875958 -GCUUCUUCUGCCACUGUUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUCUACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGUGCAAUGCAAG -((......(((....((((....((....))....))))......(((((((((((((.....))))((((((........)))))).....))))))))))))..))... ( -21.50, z-score = 0.16, R) >droVir3.scaffold_13049 20065380 111 - 25233164 GAUAGCUGUUGCUGUUGCUU-UUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUCUACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGUGCAAUGCAAC ....((.(((((.(((((((-...((....)).)))))))......(((((((((((((.....))))((((((........)))))).....))))))))))))))))... ( -30.50, z-score = -2.32, R) >droMoj3.scaffold_6680 7768637 105 - 24764193 ------GGUAGCUGUUGCUG-UUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUCUACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGAGCAAUGCAAC ------((.....((((((.-...((....))..))))))...)).(((...(((((.((((......((((((........)))))).......)))).))))).)))... ( -26.42, z-score = -1.50, R) >consensus _____________UUUGCUGAAUGCCGAAAGGAAAGCAAUUUUCCAGCAAUAUGUUGAACGUAUUAGAAUUUUGUUUACACUCGAAAUGGAAAACAUGUUGUGCAAUGCAAC ..............(((((.....((....))..)))))((((((...((((((.....))))))...((((((........))))))))))))...(((((.....))))) (-20.01 = -20.53 + 0.52)
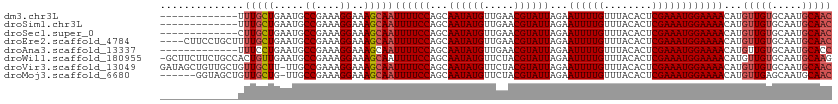

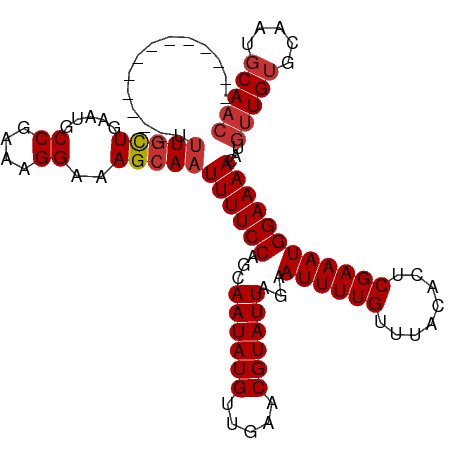

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:28:25 2011