
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 5,284,250 – 5,284,388 |
| Length | 138 |
| Max. P | 0.512867 |

| Location | 5,284,250 – 5,284,388 |
|---|---|
| Length | 138 |
| Sequences | 8 |
| Columns | 148 |
| Reading direction | forward |
| Mean pairwise identity | 61.75 |
| Shannon entropy | 0.71668 |
| G+C content | 0.40600 |
| Mean single sequence MFE | -34.13 |
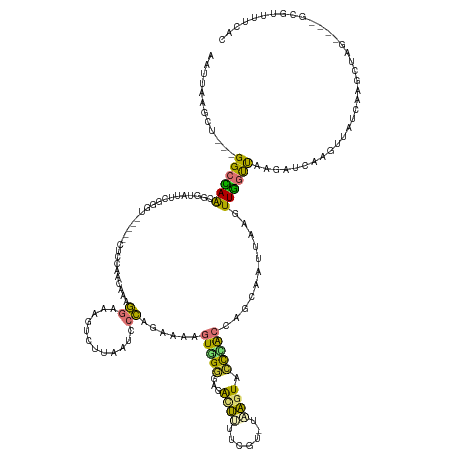
| Consensus MFE | -7.76 |
| Energy contribution | -6.58 |
| Covariance contribution | -1.18 |
| Combinations/Pair | 1.88 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.23 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.512867 |
| Prediction | RNA |
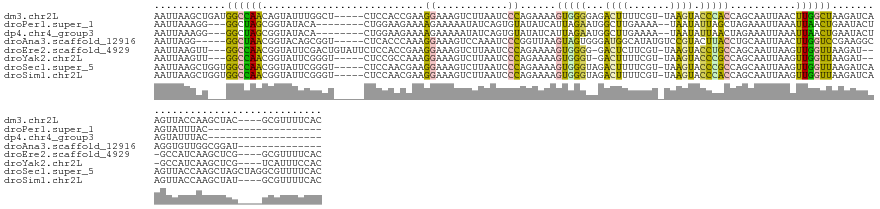
Download alignment: ClustalW | MAF
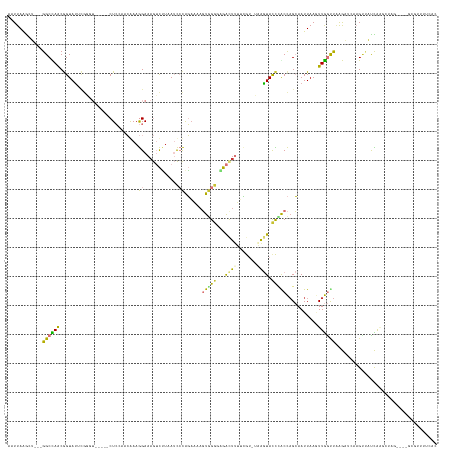
>dm3.chr2L 5284250 138 + 23011544 AAUUAAGCUGAUGGCCAACAGUAUUUGGCU-----CUCCACCGAAGGAAAGUCUUAAUCCCAGAAAAGUGGGGAGACUUUUCGU-UAAGUACCCACCAGCAAUUAACUUGGCUAAGAUCAAGUUACCAAGCUAC----GCGUUUUCAC ......((((.(((......((((((((((-----(......)).((((((((((...((((......))))))))))))))))-)))))))))).))))...((((((((......)))))))).........----.......... ( -37.80, z-score = -2.05, R) >droPer1.super_1 8541774 116 + 10282868 AAUUAAAGG---GGCUAGCGGUAUACA--------CUGGAAGAAAAGAAAAAUAUCAGUGUAUAUCAUUAGAAUGGCUUGAAAA--UAAUAUUAGCUAGAAAUUAAAUUAACUGAAUACUAGUAUUUAC------------------- ...((((..---.(((((.((((((((--------((((...............)))))))))))).......(((((......--.......)))))....................))))).)))).------------------- ( -22.58, z-score = -2.65, R) >dp4.chr4_group3 7087916 116 + 11692001 AAUUAAAGG---GGCUAGCGGUAUACA--------CUGGAAGAAAAGAAAAAUAUCAGUGUAUAUCAUUAGAAUGGCUUGAAAA--UAAUAUUAACUAGAAAUUAAAUUAACUGAAUACUAGUAUUUAC------------------- ...((((..---.(((((.((((((((--------((((...............)))))))))))).......(((.((((...--.....)))))))....................))))).)))).------------------- ( -19.36, z-score = -1.93, R) >droAna3.scaffold_12916 6303498 124 - 16180835 AAUUAGG-----GGCUAACGGUACAGCGGU-----CUCACCCAAAGGAAAGUCCAAAUCCCGGUUAAGUAGUGGGAUGGCAUAUGUCCGUACUUACCUGCAAUUAACUUGGUCCGAAGGCAGGUGUUGGCGGAU-------------- .......-----.(((((((((((.(.((.-----....)))...(((..(((((..(((((.(.....).)))))))).))...)))))))).((((((..................))))))))))))....-------------- ( -34.87, z-score = 0.58, R) >droEre2.scaffold_4929 5364546 136 + 26641161 AAUUAAGUU---GGCCAACGGUAUUCGACUGUAUUCUCCACCGAAGGAAAGUCUUAAUCCCAGAAAAGUGGGG-GACUCUUCGU-UAAGUACCUGCCAGCAAUUAAGUUGGUUAAGAU---GCCAUCAAGCUCG----GCGUUUUCAC ......(((---(((....((((((..((....(((......)))(((.(((((....((((......)))))-)))).)))))-..)))))).))))))..........((.(((((---(((.........)----))))))).)) ( -37.90, z-score = -1.44, R) >droYak2.chr2L 8410470 131 - 22324452 AAUUAAGUU---GGCCAACGGUAUUCGGGU-----CUCCGCCAAAGGAAAGUCUUAAUCCCAGAAAAGUGGGU-GACUUUUCGU-UAAGUACCCGCCAGCAAUUAAGUUGGUUAAGAU---GCCAUCAAGCUCG----UCAUUUCCAC ......(((---(((....((((((..(((-----....)))...((((((((.....((((......)))).-))))))))..-..)))))).))))))......(.((((......---)))).).......----.......... ( -34.90, z-score = -1.75, R) >droSec1.super_5 3363259 142 + 5866729 AAUUAAGCUGGUGGCCAACGGUAUUCGGGU-----CUCCAACGAAGGAAAGUCUUAAUCCCAGAAAAGUGGGUAGACUUUUCGU-UAAGUACCCGCCAGCAAUUAAGUUGGUUAAGAUCAAGUUACCAAGCUAGCUAGGCGUUUUCAC .....((((((((.....)((((...((((-----((..((((((...((((((....((((......)))).)))))))))))-).)).))))((((((......))))))...........))))..)))))))............ ( -43.10, z-score = -2.48, R) >droSim1.chr2L 5086379 138 + 22036055 AAUUAAGCUGGUGGCCAACGGUAUUCGGGU-----CUCCAACGAAGGAAAGUCUUAAUCCCAGAAAAGUGGGUAGACUUUUCGU-UAAGUACCCACCAGCAAUUAAGUUGGUUAAGAUCAAGUUACCAAGCUAU----GCGUUUUCAC .....((((((((((...........((((-----((..((((((...((((((....((((......)))).)))))))))))-).)).))))((((((......)))))).........)))))).))))..----.......... ( -42.50, z-score = -3.33, R) >consensus AAUUAAGCU___GGCCAACGGUAUUCGGGU_____CUCCAACAAAGGAAAGUCUUAAUCCCAGAAAAGUGGGGAGACUUUUCGU_UAAGUACCCACCAGCAAUUAAGUUGGUUAAGAUCAAGUUAUCAAGCUAG____GCGUUUUCAC ............((((((...........................((............))......(((((...((((.......)))).)))))...........))))))................................... ( -7.76 = -6.58 + -1.18)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:18:41 2011