
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,164,136 – 15,164,228 |
| Length | 92 |
| Max. P | 0.526811 |

| Location | 15,164,136 – 15,164,228 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 62.96 |
| Shannon entropy | 0.73312 |
| G+C content | 0.52038 |
| Mean single sequence MFE | -26.81 |
| Consensus MFE | -12.00 |
| Energy contribution | -10.61 |
| Covariance contribution | -1.39 |
| Combinations/Pair | 1.89 |
| Mean z-score | -0.51 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.526811 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
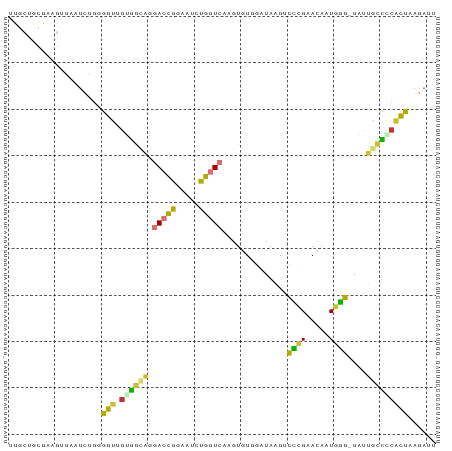
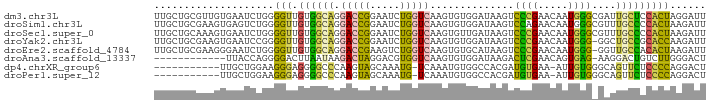
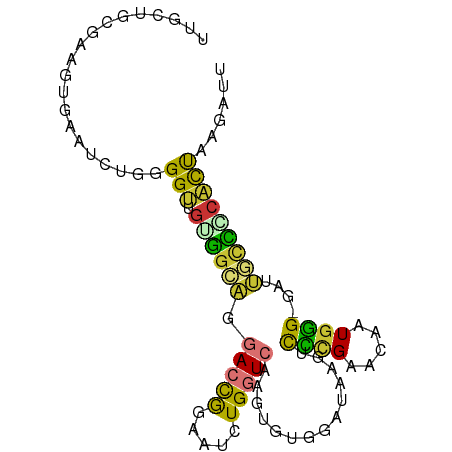
>dm3.chr3L 15164136 92 + 24543557 UUGCUGCGUUGUGAAUCUGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUGGAUAAGUCCCGAACAAUGGGCGAUUGCUCCACUAGGAUU (((((.((((((.....((((..(.(.(((.((((((...))))))...))).)...)..)))).)))))))))))....(((....))).. ( -28.10, z-score = -0.34, R) >droSim1.chr3L 14498476 92 + 22553184 UUGCUGCGAAGUGAGUCUGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUGGAUAAGUCCAGAACAAUGGGCGUUUGCCCCACUAAGAUU .........((((..((((((.((((.(((.((((((...))))))...)))..)))).)))))).....((((....))))))))...... ( -29.90, z-score = -0.98, R) >droSec1.super_0 7276794 92 + 21120651 UUGCUGCAAAGUGAAUCUGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUUGAUAAGUCCCGAACAAUGGGCGUUUGCCCCACUAAGAUU .........((((....((((..(.(((((.((((((...))))))...)))))...)..))))......((((....))))))))...... ( -26.80, z-score = -0.28, R) >droYak2.chr3L 17671566 91 + 24197627 UUGCUGCGAAGUGAAUCCGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUGGAUAAGUCCCGAACAAUGGG-GGCUGCCGCACCAAGAUU ((((..(((..((....))...)))..))))((((((...))))))..((((((...((((((........))-)))).))))))....... ( -35.60, z-score = -2.08, R) >droEre2.scaffold_4784 17425998 91 + 25762168 UUGCUGCGAAGGGAAUCUGGGGUUGUGGCAGGACCGAAGUCUGGUCAAGUGUGCAUAAGUCCCGAACAAUGGG-GGUUGCCACACUAAGAUU ((((..(((.((....))....)))..))))(((((.....))))).((((((..(((.(((((.....))))-).))).))))))...... ( -28.80, z-score = -0.69, R) >droAna3.scaffold_13337 5468248 79 - 23293914 ------------UUACCAGGGGACUUAAUAAGACUAGGACGUGGUCAAGUGUGGAUAAGACUCGAACAGUGAG-AAGGACUGUCUUGGGACU ------------...((((....).......(((((.....))))).....)))......((((((((((...-....))))).)))))... ( -16.70, z-score = 0.51, R) >dp4.chrXR_group6 12741728 79 - 13314419 -----------UUGCUGGAAGGGAGGGGCCCAAGUAGCAAAUG-UCAAAUGUGGCCACGAUGUGAA-AUUGUGGGCAGUUCUCCCCAGGACU -----------((((((...(((.....)))...))))))...-.......((.(((((((.....-))))))).))(((((....))))). ( -24.30, z-score = -0.13, R) >droPer1.super_12 1159853 79 - 2414086 -----------UUGCUGGAAGGGAGGGGCCCAAGUAGCAAAUG-UCAAAUGUGGCCACGAUGUGAA-AUUGUGGGCAGUUCUCCCCAGGACU -----------((((((...(((.....)))...))))))...-.......((.(((((((.....-))))))).))(((((....))))). ( -24.30, z-score = -0.13, R) >consensus UUGCUGCGAAGUGAAUCUGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUGGAUAAGUCCCGAACAAUGGG_GAUUGCCCCACUAAGAUU ....................(((.((((((.(((((.....)))))..............((((.....))))....)))))))))...... (-12.00 = -10.61 + -1.39)


| Location | 15,164,136 – 15,164,228 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 62.96 |
| Shannon entropy | 0.73312 |
| G+C content | 0.52038 |
| Mean single sequence MFE | -26.81 |
| Consensus MFE | -12.00 |
| Energy contribution | -10.61 |
| Covariance contribution | -1.39 |
| Combinations/Pair | 1.89 |
| Mean z-score | -0.51 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.526811 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 15164136 92 + 24543557 UUGCUGCGUUGUGAAUCUGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUGGAUAAGUCCCGAACAAUGGGCGAUUGCUCCACUAGGAUU (((((.((((((.....((((..(.(.(((.((((((...))))))...))).)...)..)))).)))))))))))....(((....))).. ( -28.10, z-score = -0.34, R) >droSim1.chr3L 14498476 92 + 22553184 UUGCUGCGAAGUGAGUCUGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUGGAUAAGUCCAGAACAAUGGGCGUUUGCCCCACUAAGAUU .........((((..((((((.((((.(((.((((((...))))))...)))..)))).)))))).....((((....))))))))...... ( -29.90, z-score = -0.98, R) >droSec1.super_0 7276794 92 + 21120651 UUGCUGCAAAGUGAAUCUGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUUGAUAAGUCCCGAACAAUGGGCGUUUGCCCCACUAAGAUU .........((((....((((..(.(((((.((((((...))))))...)))))...)..))))......((((....))))))))...... ( -26.80, z-score = -0.28, R) >droYak2.chr3L 17671566 91 + 24197627 UUGCUGCGAAGUGAAUCCGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUGGAUAAGUCCCGAACAAUGGG-GGCUGCCGCACCAAGAUU ((((..(((..((....))...)))..))))((((((...))))))..((((((...((((((........))-)))).))))))....... ( -35.60, z-score = -2.08, R) >droEre2.scaffold_4784 17425998 91 + 25762168 UUGCUGCGAAGGGAAUCUGGGGUUGUGGCAGGACCGAAGUCUGGUCAAGUGUGCAUAAGUCCCGAACAAUGGG-GGUUGCCACACUAAGAUU ((((..(((.((....))....)))..))))(((((.....))))).((((((..(((.(((((.....))))-).))).))))))...... ( -28.80, z-score = -0.69, R) >droAna3.scaffold_13337 5468248 79 - 23293914 ------------UUACCAGGGGACUUAAUAAGACUAGGACGUGGUCAAGUGUGGAUAAGACUCGAACAGUGAG-AAGGACUGUCUUGGGACU ------------...((((....).......(((((.....))))).....)))......((((((((((...-....))))).)))))... ( -16.70, z-score = 0.51, R) >dp4.chrXR_group6 12741728 79 - 13314419 -----------UUGCUGGAAGGGAGGGGCCCAAGUAGCAAAUG-UCAAAUGUGGCCACGAUGUGAA-AUUGUGGGCAGUUCUCCCCAGGACU -----------((((((...(((.....)))...))))))...-.......((.(((((((.....-))))))).))(((((....))))). ( -24.30, z-score = -0.13, R) >droPer1.super_12 1159853 79 - 2414086 -----------UUGCUGGAAGGGAGGGGCCCAAGUAGCAAAUG-UCAAAUGUGGCCACGAUGUGAA-AUUGUGGGCAGUUCUCCCCAGGACU -----------((((((...(((.....)))...))))))...-.......((.(((((((.....-))))))).))(((((....))))). ( -24.30, z-score = -0.13, R) >consensus UUGCUGCGAAGUGAAUCUGGGGUUGUGGCAGGACCGGAAUCUGGUCAAGUGUGGAUAAGUCCCGAACAAUGGG_GAUUGCCCCACUAAGAUU ....................(((.((((((.(((((.....)))))..............((((.....))))....)))))))))...... (-12.00 = -10.61 + -1.39)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:28:04 2011