
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,139,285 – 15,139,341 |
| Length | 56 |
| Max. P | 0.980893 |

| Location | 15,139,285 – 15,139,341 |
|---|---|
| Length | 56 |
| Sequences | 3 |
| Columns | 60 |
| Reading direction | forward |
| Mean pairwise identity | 85.00 |
| Shannon entropy | 0.21008 |
| G+C content | 0.33955 |
| Mean single sequence MFE | -12.70 |
| Consensus MFE | -11.29 |
| Energy contribution | -11.73 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.89 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.857081 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
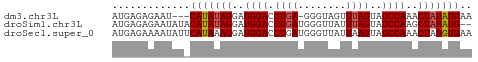
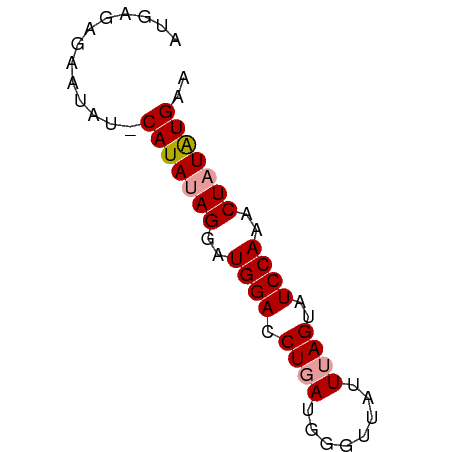
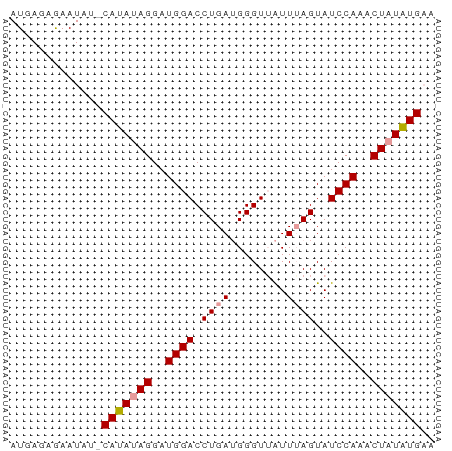
>dm3.chr3L 15139285 56 + 24543557 AUGAGAGAAU---CAUAUAGGAUGGACCUGA-GGGUAGUUUAGUAUCCAAACUAUAUGAA .........(---(((((((..((((.((((-(.....)))))..))))..)))))))). ( -16.10, z-score = -3.00, R) >droSim1.chr3L 14473606 58 + 22553184 AUGAGAGAAUAUACAUAUAGGAUGGACCUGAUGGGUUAUUUAGUAUCCAAGCUAUAUG-- .............(((((((..((((.((((........))))..))))..)))))))-- ( -12.10, z-score = -1.19, R) >droSec1.super_0 7252269 60 + 21120651 AUGAGAAAAUAUUCAUAAAGGAUGGACCUGAUGGGUUAUUAAGUAUCCAAACUAUGUGAA ...........((((((.((..((((((....))...........))))..)).)))))) ( -9.90, z-score = -0.50, R) >consensus AUGAGAGAAUAU_CAUAUAGGAUGGACCUGAUGGGUUAUUUAGUAUCCAAACUAUAUGAA .............(((((((..((((.((((........))))..))))..))))))).. (-11.29 = -11.73 + 0.45)
| Location | 15,139,285 – 15,139,341 |
|---|---|
| Length | 56 |
| Sequences | 3 |
| Columns | 60 |
| Reading direction | reverse |
| Mean pairwise identity | 85.00 |
| Shannon entropy | 0.21008 |
| G+C content | 0.33955 |
| Mean single sequence MFE | -11.35 |
| Consensus MFE | -9.96 |
| Energy contribution | -10.63 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.88 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.980893 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
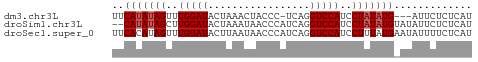
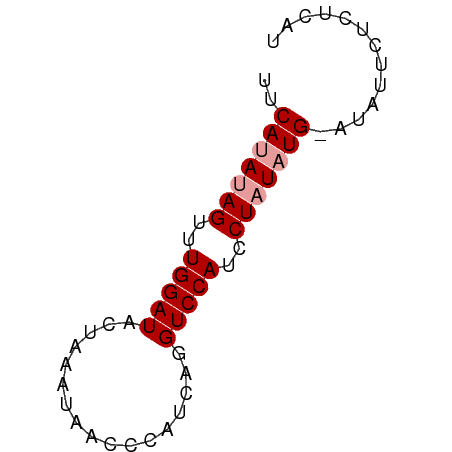
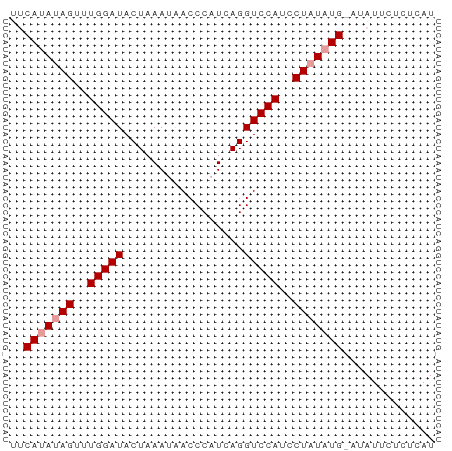
>dm3.chr3L 15139285 56 - 24543557 UUCAUAUAGUUUGGAUACUAAACUACCC-UCAGGUCCAUCCUAUAUG---AUUCUCUCAU .((((((((..(((((............-....)))))..)))))))---)......... ( -14.69, z-score = -3.80, R) >droSim1.chr3L 14473606 58 - 22553184 --CAUAUAGCUUGGAUACUAAAUAACCCAUCAGGUCCAUCCUAUAUGUAUAUUCUCUCAU --(((((((..(((((.................)))))..)))))))............. ( -11.53, z-score = -2.40, R) >droSec1.super_0 7252269 60 - 21120651 UUCACAUAGUUUGGAUACUUAAUAACCCAUCAGGUCCAUCCUUUAUGAAUAUUUUCUCAU ((((...((..(((((.................)))))..))...))))........... ( -7.83, z-score = -0.85, R) >consensus UUCAUAUAGUUUGGAUACUAAAUAACCCAUCAGGUCCAUCCUAUAUG_AUAUUCUCUCAU ..(((((((..(((((.................)))))..)))))))............. ( -9.96 = -10.63 + 0.67)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:28:02 2011